Entry Clone Source: Origene |
Entry Clone Accession: NM_003829 Variant |
SGC Construct ID: MPDZA-c119 |
GenBank GI number: gi|4505231 |
Vector: pNIC28-BSA4. Details [ PDF ]; Sequence [ FASTA ] or [ GenBank ] |
Tags and additions: N-terminal hexahistidine tag before TEV cleavage site C-terminal PDZ recognition motif |
Sequence:
mhhhhhhssgvdlgtenlyfqsmQPRRVE
LWREPSKSLGISIVGGRGMGSRLSNGEVM
RGIFIKHVLEDSPAGKNGTLKPGDRIVEV
DGMDLRDASHEQAVEAIRKAGNPVVFMVQ
SIISTRL
Lower case characters highlight the N-terminal hexahistidine tag and TEV cleavage recognition site
STRL: C-terminal extension containing PDZ recognition motif |
Host : BL-21(DE3)R3 phage resistant |
Growth medium, induction protocol: Transformed 50 µl competent BL-21 (DE3) phage resistant cells with 10 µl of the plasmid DNA and plated out onto LB plate plus 50 µg/ml kanamycin. The next day colonies were picked out into fresh deep well blocks containing 1 ml TB + 50 µg/ml kanamycin. These were grown overnight and glycerol stocks prepared by adding 333 µl of 60 % glycerol to 1 ml of cell suspension, mixing and then storing in a -80°C freezer.
The glycerol stock was used to innoculate 20 mls of TB + 50 µg/ml kanamycin which was grown overnight at 37°C as a starter culture for a 1 litre growth. The large scale growth was grown at 37°C until approximately 30 mins before induction when the temperature was lowered to 25°C. Protein production was induced with the addition of 1mM IPTG. The next day cells were harvested by centrifugation at 4000 rpm for 15 minutes. The pellet was then stored in the -80°C freezer. |
Extraction buffer, extraction method: Lysis buffer: 10mM Imidazole, 300mM NaCl, 50mM pH8.0 NaH2PO4, 0.5mM TCEP, 1x complete PI EDTA free tablet/50mls.
The pellet (19.52 gms) was resuspended with 3x volume of lysis buffer (approximately 50 mls final) by intermitently placing the pellet in a 37°C water bath and vortexing. Once resuspended the cells were (1) broken by one passage through the Constant Systems cell breaker; (2) sonicating; (3) DNA precipitation with the addition of PEI to a final concentration of 0.15 % for 30 mins on ice followed by a 17,000 rpm at 4°C to remove precipitation; (4) the supernatant was filtered through a GF/0.2 µM serum acrodiscs. |
Column 1 : Ni-affinity, HisTrap, 1 ml (GE/Amersham) |
Buffers: Affinity binding buffer: 10mM Imidazole, 300mM NaCl, 50mM pH8.0 NaH2PO4, 0.5mM TCEP. Affinity wash buffer: 50mM Imidazole, 300mM NaCl, 50mM pH8.0 NaH2PO4, 0.5mM TCEP. Affinity Elution Buffer: 250mM Imidazole, 300mM NaCl, 50mM pH8.0 NaH2PO4, 0.5mM TCEP |
Procedure: The cell extract was loaded on the column at 0.8 ml/min on an AKTA-express system (GE/Amersham). The column was then washed with 10 column volumes of Affinity Binding buffer, 10 column volumes of Affinity wash buffer, and then eluted with Affinity elution buffer at 0.8 ml/min. The eluted peak of A280 was automatically collected. |
Column 2 : Gel filtration, Hiload 16/60, S75 16/60 - 120 ml |
Buffers : Gel Filtration: 10mM pH7.4 Hepes, 500mM NaCl, 5% glycerol, 0.5mM TCEP |
Procedure: The eluted fractions from the Ni-affinity Histrap columns were loaded on the gel filtration column in GF buffer at 1.0 ml/min. Eluted proteins were collected in 1 ml fractions. |
Concentration : Using a Centricon 10 K cutoff concentrator the MPDZA-p065 pooled fractions was concentrated to 57.6 mg/ml. Concentration was determined from the absorbance at 280 nm. |
Mass spec characterization : Expected mass spect for full length construct 13590.4; recorded mass 13590.7. |
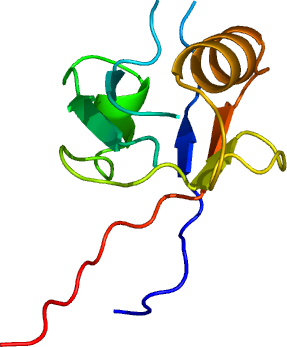
Crystallisation: 2CFC: Crystals grew from a 1:2 ratio mix of MPDZA-to-reservoir ( 30% mPEG 2K; 0.10M KSCN ); 2IWQ: Crystals grew from a 1:2 ratio mix of MPDZA-to-reservoir (0.2 M NaF; 0.1M BTProp pH 8.5; 20 % PEG 3350; 10 % ethylene glycol). |
Data Collection: Resolution: 1.76Å (2CFC) and 1.8Å (2IWQ); X-ray source: Synchrotron SLS -X10, single wavelength. |