Vector: pNIC-Zb |
Amplified construct sequence:
ATGCACCATCATCATCATCATTCTTC
TGGTGTGGATAACAAGTTCAACAAGG
AGCGTCGAAGAGCTCGCCGTGAAATT
CGCCATCTGCCGAACCTGAACCGCGA
ACAGCGTCGCGCATTTATTCGCAGCC
TGCGCGATGATCCGAGCCAGAGCGCG
AACCTGCTGGCGGAAGCGAAGAAGCT
GAACGATGCGCAGCCGAAGGGTACCG
AGAACCTGTACTTCCAATCCATGGGG
CCGGAGGATGAGCTGCCCGACTGGGC
CGCCGCCAAAGAGTTTTACCAGAAGT
ACGACCCTAAGGACGTCATCGGCAGA
GGAGTGAGCTCTGTGGTCCGCCGTTG
TGTTCATCGAGCTACTGGCCACGAGT
TTGCGGTGAAGATTATGGAAGTGACA
GCTGAGCGGCTGAGTCCTGAGCAGCT
GGAGGAGGTGCGGGAAGCCACACGGC
GAGAGACACACATCCTTCGCCAGGTC
GCCGGCCACCCCCACATCATCACCCT
CATCGATTCCTACGAGTCTTCTAGCT
TCATGTTCCTGGTGTTTGACCTGATG
CGGAAGGGAGAGCTGTTTGACTATCT
CACAGAGAAGGTGGCCCTCTCTGAAA
AGGAAACCAGGTCCATCATGCGGTCT
CTGCTGGAAGCAGTGAGCTTTCTCCA
TGCCAACAACATTGTGCATCGAGATC
TGAAGCCCGAGAATATTCTCCTAGAT
GACAATATGCAGATCCGACTTTCAGA
TTTCGGGTTCTCCTGCCACTTGGAAC
CTGGCGAGAAGCTTCGAGAGTTGTGT
GGGACCCCAGGGTATCTAGCGCCAGA
GATCCTTAAATGCTCCATGGATGAAA
CCCACCCAGGCTATGGCAAGGAGGTC
GACCTCTGGGCCTGTGGGGTGATCTT
GTTCACACTCCTGGCTGGCTCGCCAC
CCTTCTGGCACCGGCGGCAGATCCTG
ATGTTACGCATGATCATGGAGGGCCA
GTACCAGTTCAGTTCCCCCGAGTGGG
ATGACCGTTCCAGCACTGTCAAAGAC
CTGATCTCCAGGCTGCTGCAGGTGGA
TCCTGAGGCACGCCTGACAGCTGAGC
AGGCCCTACAGCACCCCTTCTTTGAG
CGTTGA
|
Final protein sequence (Tag sequence in lowercase):
smGPEDELPDWAAAKEFYQKYDPKDV
IGRGVSSVVRRCVHRATGHEFAVKIM
EVTAERLSPEQLEEVREATRRETHIL
RQVAGHPHIITLIDSYESSSFMFLVF
DLMRKGELFDYLTEKVALSEKETRSI
MRSLLEAVSFLHANNIVHRDLKPENI
LLDDNMQIRLSDFGFSCHLEPGEKLR
ELCGTPGYLAPEILKCSMDETHPGYG
KEVDLWACGVILFTLLAGSPPFWHRR
QILMLRMIMEGQYQFSSPEWDDRSST
VKDLISRLLQVDPEARLTAEQALQHP
FFER
(Gly6 to Arg293)
The N-terminal residues, sm, derive from the vector following TEV protease digestion to remove the expression tag. |
Tags and additions: N-terminal, TEV cleavable hexahistidine tag and Zbasic tag |
Host: BL21 (DE3)R3-pRARE2 (Phage resistant strain).
|
Growth medium, induction protocol: The construct DNA was transformed into homemade chemically competent cells of the expression strain by a standard heat shock procedure. A number of colonies from the transformation were used to inoculate 1ml of LB media containing 50µg/ml kanamycin and 34µg/ml chloramphenicol, which was placed in a 37°C shaker overnight. The next day glycerol stocks were prepared from this overnight culture. A glycerol stock was used to inoculate 40ml of LB media containing 50µg/ml kanamycin and 34µg/ml chloramphenicol, which was placed in a 37°C shaker overnight. The next day 2x10ml of this starter culture was used to inoculate 2x1L of LB media containing 50µg/ml kanamycin in 2x2L baffled shaker flasks. When the OD600 was approximately 0.5, the temperature was reduced to 20°C and the cells were induced by the addition of 0.5mM IPTG. The expression was continued overnight. Cells were spun at 5500rpm for 10 mins and the pellets resuspended in Lysis Buffer and then frozen at -20°C.
Lysis buffer: 50 mM HEPES, pH 7.5; 500 mM NaCl; 5% Glycerol; 5 mM Imidazole; 0.5 mM TCEP; 0.5 mM PMSF.
Extraction buffer, extraction method: The resuspended cell pellet was thawed and lysed by sonication. The sample was diluted to 90ml volume and PEI (polyethyleneimine) was added to a final concentration of 0.15%. The cell debris and precipitated DNA were spun down (17000rpm, JA17 rotor, 45 min). |
Column 1: 3ml of Ni-NTA in a 1cm diameter gravity flow column. |
Column 1 Buffers:
Binding Buffer:
50 mM HEPES, pH 7.5; 500 mM NaCl; 5% glycerol; 5 mM Imidazole; 0.5 mM TCEP.
Wash Buffer: 50 mM HEPES, pH 7.5; 500 mM NaCl; 5% glycerol; 25 mM Imidazole; 0.5 mM TCEP.
Elution Buffer: 50 mM HEPES, pH 7.5; 500 mM NaCl; 5% glycerol; 250 mM Imidazole; 0.5 mM TCEP. |
Column 1 Procedure: The clarified supernatant was passed through the column. The column was washed with 60ml of binding Buffer and 50ml of Wash Buffer. 15ml of Elution Buffer was passed through to elute the protein. |
Column 2: 5ml HiTrap S |
Column 2 Buffers:
Buffer A: 25 mM HEPES, pH 7.0; 200 mM NaCl; 0.5 mM TCEP.
Buffer B: 25 mM HEPES, pH 7.0; 2 mM NaCl; 0.5 mM TCEP. |
Column 2 Procedure: The eluted protein was diluted with 25ml HEPES pH 7.0 to a NaCl concentration of approximately 200 mM. The protein was loaded onto a 5ml HiTrap S column at 4ml/min (pre-equilibrated in Buffer A), and eluted with a gradient to Buffer B. |
Enzymatic treatment: The fractions containing PHKG2A were pooled and TEV protease was added. The sample was left at 4°C overnight. |
Column 3: S200 16/60 Gel Filtration |
Column 2 Buffer: 25 mM HEPES, pH 7.4; 500 mM NaCl; 0.5 mM TCEP. |
Column 2 Procedure: The protein was concentrated to 5ml volume and injected onto an S200 16/60 GF column (pre-equilibrated in GF Buffer) at 1.0 ml/min. 1.75ml fractions were collected. |
Protein concentration: The pooled fractions from column 3 were concentrated to 4.5mg/ml (measured by 280nm absorbance). |
Mass spectrometry characterization:
Measured: 33394.8
Expected: 33394.3
|
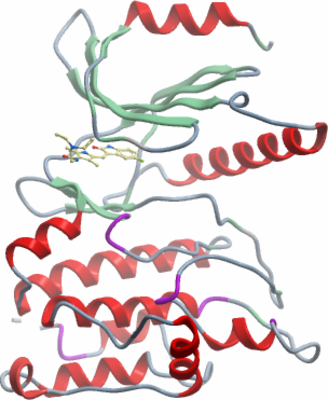
Crystallisation: Crystals grew from a 2:1 ratio of protein and precipitant solution (25% PEG3350, 0.1M BisTris pH 5.5), using the vapour diffusion method. |
Data collection: Crystals were cryo-protected by equilibration into precipitant solution containing 25% ethylene glycol, and then flash frozen in liquid nitrogen. Data was collected at Diamond beamline I03. |