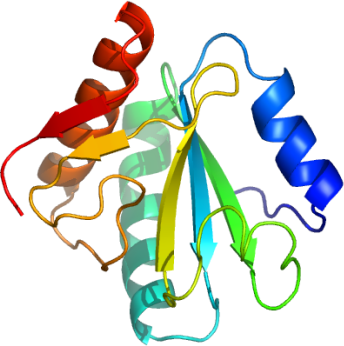
The crystal structure of U5 small-nuclear-ribonucleoprotein-particle-(snRNP)-specific
15 kD protein (U5-15kD) has been solved for Plasmodium yoelii
(PY07357).
It has 85% identity to its Plasmodium falciparum ortholog
PFL1520w
which is annotated as a putative DIM1 protein homolog.
The structure of U5-15kD is similar to thioredoxin,
hence it has also been dubbed thioredoxin-like protein 4A,
with 37 amino acids distinguishing the two structures and giving rise to changes
that likely support binding sites to other spiceosomal proteins or RNA.
The supramolecular spliceosome catalyzes the removal of introns from nuclear pre-mRNA
which involves two sequential transesterification reactions.
The spliceosome is comprised of more than 80 proteins,
including the small nuclear ribonucleoprotein particles (snRNPs) U1, U2, U4/U6, and U5,
as well as non-snRNP splicing factors.
Each snRNP contains either one of U1, U2, U5 or two U4/U6 snRNPS,
plus some of other integral proteins mentioned.
Biochemically, U5 is the most complex, including seven core snRNPs and nine U5 snRNP-specific proteins,
which are involved directly in splicing.
These include the 220 kDa protein which is likely at the catalytic centre;
the ATPases U5-100 kD and U5-200 kD proteins, which are involved in complexes with RNA unwindase;
and the GTPase U5-116 kD with homology to ribosomal translocase.
Like its human ortholog, Py-U5-15kD exhibits a thioredoxin-like fold characterized
by its pairs of parallel and anti-parallel four-stranded beta-sheet and flanked by three alpha-helices.
Py-U5-15kD is 75% identical to is human ortholog (1QGV);
however, there are an additional 37 residues in the U5-15kDs over the human thioredoxin,
which accounts for the additional parallel C-terminal beta-strand leading to a distortion
of the structure with respect to the human thioredoxin structure (1ERU).
Another feature distinguishing the thioredoxin structure from the U5-15kD structures
is the presence of a disulfide bond.
The thioredoxins, as protein disulfide isomerases (PDI),
have a Cys-X-X-Cys structural motif which is a functional disulfide
located at the N-terminus of alpha-helix-2.
In the human ortholog, however, this is replaced by Asp-X-X-Cys (Asp35 and Cys38),
identical to the Plasmodium yoelii ortholog (Asp53 and Cys56 for numbering with His-tag).
Even though there is no Cys35 to form a disulfide with Cys38 in the human U5-15kD,
Cys38 forms a disulfide with Cys79, close to the N-terminus of strand beta-3.
The authors of the human U5-15kD structure note that in the electron density map,
there is also a less occupied alternative conformation of the Cys79 side-chain
indicative of a mixture of oxidized and reduced forms in the crystal.1
For the Plasmodium yoelii ortholog, however, there is no Cys79 (human numbering),
as it is replaced with a Val (Val 97 with Py-U5-15kD numbering including His-tag).
There are two unconserved Cys at positions Cys45 and Cys74
that are potentially within disulfide forming distance, but they exist as a dithiol motif.
In the human ortholog, the authors speculated that U5-15kD may have disulfide isomerase activity because of the observation of a disulfide between Cys39 and Cys79, but this was not observed.1
The authors concluded that this was consistent with the observation that only 5 out of the 12 U5-kD sequences available had a Cys at position 79.