Entry Clone Source: TKC |
Entry Clone Accession: n/a |
SGC Construct ID: PIM1A-c001 |
GenBank GI number: gi|4505811 |
Entry clone accession : The expressed protein has the sequence from gi|33304198 which has the isoform change R250G compared to gi|4505811. |
Expressed protein sequence:
mhhhhhhssgvdlgtenlyfqsMLLSKIN
SLAHLRAAPCNDLHATKLAPGKEKEPLES
QYQVGPLLGSGGFGSVYSGIRVSDNLPVA
IKHVEKDRISDWGELPNGTRVPMEVVLLK
KVSSGFSGVIRLLDWFERPDSFVLILERP
EPVQDLFDFITERGALQEELARSFFWQVL
EAVRHCHNCGVLHRDIKDENILIDLNRGE
LKLIDFGSGALLKDTVYTDFDGTRVYSPP
EWIRYHRYHGRSAAVWSLGILLYDMVCGD
IPFEHDEEIIGGQVFFRQRVSSECQHLIR
WCLALRPSDRPTFEEIQNHPWMQDVLLPQ
ETAEIHLHSLSPGPS |
Vector: pLIC- SGC |
Tags and additions: mhhhhhhssgvdlgtenlyfq*s(m). N-terminal his6 tag, TEV-protease cleavable (*) |
Host : BL21(DE3) |
Growth medium, induction protocol: 1 ml from a 50 ml overnight culture in LB, 100µg/ml ampicillin was used to inoculate 1 litre of LB medium containing 100µg/ml ampicillin. Cultures were grown at 37°C until they reached an OD 600 of 0.3 and then cooled to 18°C. Expression was induced for 4 hours using 1 mM IPTG at an OD 600 of 0.6. The cells were collected by centrifugation, transferred in 30 ml binding buffer to 50 ml tubes, and frozen at -20°C. |
Binding buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 5% Glycerol. |
Extraction buffer, extraction method: The frozen cells were thawed on ice. Binding buffer (plus 1 mM PMSF) was added to a final volume of 50 ml. Cells were lysed using a high pressure cell disruptor. The lysate was centrifuged at 17,000 RPM for 30 minutes and the supernatant collected for purification. |
Column 1 : DEAE cellulose (DE52, Whatmann), 10 g of resin in 2.5 x 20 cm column |
Buffers: Binding buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 5% Glycerol |
Procedure: Ion exchange - Nucleic acid removal.
The resin was hydrated in 2.5M NaCl, then washed with 50 ml binding buffer prior to loading the sample. Supernatant was applied at gravity flow, followed by a wash with 50 ml binding buffer. The column flow-through was collected. |
Column 2 : Ni-NTA (Qiagen), 5 ml of 50% slurry in 1.5 x 10 cm column |
Buffers : Binding buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 5% Glycerol; Wash buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 20 mM Imidazole, 5% glycerol; Elution buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 50 to 250 mM Imidazole, 5% Glycerol. |
Procedure: The flowthrough from column 1 was loaded by gravity flow on the Ni-NTA column. The column was then washed with 50 ml wash buffer under gravity flow. The protein was eluted by applying 5-ml portions of elution buffer with increasing concentration of imidazole (50 mM, 100 mM, 150 and 250 mM); fractions were collected until essentially all protein was eluted. After elution DTT was added to a final concentration of 10 mM. |
Column 3: Ion exchange Mono Q column |
Buffers: A : 50 mM Hepes pH 7.5; B : 50 mM Hepes pH 7.5, 2 M NaCl; |
Procedure: Ion exchange was performed after overnight enzymatic treatment at 4°C (see below). Partially dephosphorylated PIM1 was applied to a 1 ml MonoQ column in buffer A and eluted from the column by a linear gradient with buffer B. Non-phosphorylated PIM1 eluted at ~250 mM NaCl and was resolved from some residual singly phosphorylated PIM1 (at Ser261) which eluted at ~280 mM NaCl. |
Concentration : PIM1 samples containing either unphosphorylated or singly phosphorylated protein were pooled and concentrated in Centricons (10 kDa cut off) to 10 mg/ml. |
Enzymatic treatment : (Dephosphorylation and Histag cleavage) Prior to monoQ, samples containing Ni-NTA purified PIM1 were pooled and 20µg GST-lambda phosphatase and 20µg TEV protease added for overnight incubation at 4°C. Protein solution contained 10 mM DTT and 0.05 mM MnCl 2 (higher MnCl 2 concentrations caused precipitation). |
Mass spec characterization : Masses of purified proteins were confirmed by LC-MS, using an Agilent LC/MSD TOF system with reversed-phase HPLC coupled to electrospray ionisation and an orthogonal time-of-flight mass analyser. Proteins were desalted prior to mass spectrometry by rapid elution off a C3 column with a gradient of 5-95% acetonitrile in water with 0.1% formic acid. The purified PIM1 protein was homogeneous and had an experimental mass of 35546 Da (unphosphorylated fraction) or 35626 (Ser261, singly phosphorylated fraction) as expected from its primary structure |
Crystallisation:
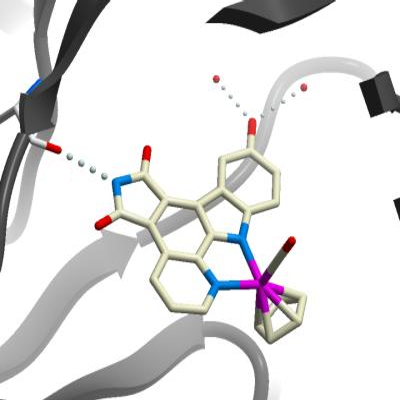
R - 5 Complex: Non-phosphorylated PIM1 crystals were grown at 4°C in 1.8µL sitting drops, mixing 1.2µL PIM1 (10 mg/mL in 50mM HEPES pH 7.5, 250 mM NaCl, 5 % glycerol, 10 mM DTT) with 0.6µL precipitant stock (0.12 M sodium citrate, 60 mM BisTrisPropane pH 6.5, 12% PEG 3350, 6% ethylene glycol, and 0.3% DMSO). The ruthenium half sandwich complex ( R )- 5 was added to the protein prior to concentration from a 10 mM stock in DMSO to give a final concentration of 1 mM. ( S )- 6 complex: Singly phosphorylated PIM1 (ser261) crystals were grown at 4°C in 1.8µL sitting drops, mixing 1.2µL PIM1 (10 mg/mL in 50mM HEPES pH 7.5, 250 mM NaCl, 5 % glycerol, 10 mM DTT) with 0.6µL precipitant stock (0.12 M potassium thiocyanate, 60 mM BisTrisPropane pH 7.5, 12% PEG 3350, 6% ethylene glycol, and 0.3% DMSO). The ruthenium half sandwich complex ( S )- 6 was added to the protein prior to concentration from a 10 mM stock in DMSO to give a final concentration of 1 mM. EA72E2 Complex: Non-phosphorylated PIM1 crystals were grown at 4°C in 150 nL sitting drops, mixing 100 nL PIM1 (10 mg/mL in 50mM HEPES pH 7.5, 250 mM NaCl, 5 % glycerol, 10 mM DTT) with 50 nL precipitant stock ( 10% PEG 10K, 0.2M MgCl 2 , 0.1M Tris pH 7.0). The ruthenium half sandwich complex EA72E2 was added to the protein prior to concentration from a 2 mM stock in DMSO to give a final concentration of 1 mM. |
Data Collection/Resolution:
R - 5 Complex (PDB 2 BZH ): 1.9Å , X-ray source: rotating anode (Rigaku FR-E SuperBright), single wavelength
( S )- 6 complex (PDB 2BZI): 1.9Å , X-ray source: Berlin synchrotron BESSY beamline BL2
EA72E2 Complex (PDB 2BZJ): 2.05Å , X-ray source: Swiss Light Source synchrotron beamline SLS -X10. |