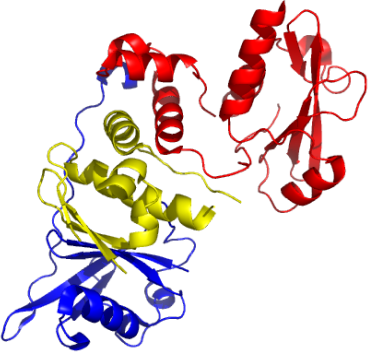
SOCS2-ElonginC-ElonginB complex
PDB:2C9W
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:gi|4507263
Entry Clone Source:MGC
SGC Clone Accession:Tag:mhhhhhhssgvdlgtenlyfq*s(m); N-terminal his6 tag, TEV-protease cleavable (*)
Host:BL21(DE3)
Construct
Prelude:Sequence:SOCS2:
mhhhhhhssgvdlgtenlyfqsmQAARLAKALRELGQTGWYWGSMTVNEAKEKLKEAPEGTFLIRDSSHSDYLLTISVKTSAGPTNLRIEYQDGKFRLDSIICVKSKLKQFDSVVH LIDYYVQMCKDKRTGPEAPRNGTVHLYLTKPLYTSAPSLQHLCRLTINKCTGAIWGLPLPTRLKDYLEEYKFQV
ElonginC:
MMYVKLISSDGHEFIVKREHALTSGTIKAMLSGPGQFAENETNEVNFREIPSHVLSKVCMYFTYKVRYTNSSTEIPEFPIAPEIALELLMAANFLDC
ElonginB:
MDVFLMIRRHKTTIFTDAKESSTVFELKRIVEGILKRPPDEQRLYKDDQLLDDGKTLGECGFTSQTARPQAPATVGLAFRADDTFEALCIEPFSSPPELPDVMKPQDSGSSANEQAVQ
Vector:pLIC-SGC (SOCS2), pACYCDUET-1 (ElonginBC)
Growth
Medium:Antibiotics:Procedure:Purification
BuffersProcedureColumn 1: Ion exchange - Nucleic acid removal - DEAE cellulose (DE52, Whatmann), 25 g of resin in 2.5 x 20 cm column. The resin was hydrated in 2.5M NaCl, then washed with 20 ml binding buffer prior to loading the sample.
Buffers: Binding buffer
Procedure: Supernatant was applied at gravity flow, followed by a wash with 50 ml binding buffer. The column flow-through was collected.
Column 2: Ni-affinity - Ni-NTA (Qiagen), 5 ml of 50% slurry in 1.5 x 10 cm column, washed with binding buffer.
Buffers : Binding buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 5% Glycerol; Wash buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 20 mM Imidazole, 5% glycerol; Elution buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 50 to 250 mM Imidazole , 5% Glycerol
Procedure: The flowthrough from column 1 was loaded by gravity flow on the Ni-NTA column. The column was then washed with 8 x 10 ml wash buffer at gravity flow. The protein was eluted by gravity flow by applying 5-ml portions of elution buffer with increasing concentration of imidazole (50 mM, 100 mM, 150 and 250 mM); fractions were collected until essentially all protein was eluted. After elution DTT was added to a final concentration of 10 mM.
Enzymatic treatment : (Histag cleavage) Samples containing the SOCS2-ElonginC/B complex were pooled and 20 µg TEV protease added for overnight incubation at 4°C.
Column 3: HiLoad S75 Gel Filtration
Procedure: Protein was ran in 25 mM HEPES pH 7.5, 250 mM NaCl and the ternary complex fractions pooled.
Column 4: Ion exchange Mono Q column .
Buffers: A: 50 mM Hepes pH 7.5; B: 50 mM Hepes pH 7.5, 2M NaCl
Procedure: SOCS2-ElonginC/B complex was applied to a 1 ml MonoQ column in buffer A and eluted from the column by a linear gradient with buffer B.
Concentration : Samples containing the ternary complex were pooled and concentrated in Centricons (10 kDa cut off) to 22 mg/ml. Protein composition was confirmed us ing LC-ESI MS-Tof.
Extraction
BuffersProcedureThe frozen cells were thawed on ice and binding buffer (plus 1 mM PMSF, 0.5 mM TCEP) added to a final volume of 225 ml. Cells were lysed using a high pressure cell disruptor. The lysate was centrifuged at 17,000 RPM for 30 minutes and the supernatant collected for purification.
Concentration:LigandMassSpec:Masses of purified proteins were confirmed by LC-MS, using an Agilent LC/MSD TOF system with reversed-phase HPLC coupled to electrospray ionisation and an orthogonal time-of-flight mass analyser. Proteins were desalted prior to mass spectrometry by rapid elution off a C3 column with a gradient of 5-95% acetonitrile in water with 0.1% formic acid.
Crystallization:Crystals were obtained at 4°C in 1 µl sitting drops using a precipitant of 0.08 M Na-Cacodylate pH 6.5, 0.16 M NaCl, 1.6 M (NH4)2SO4.
NMR Spectroscopy:Data Collection:Data were collected at the SLS. The structure was solved by molecular replacement using 1LM8.
Data Processing: