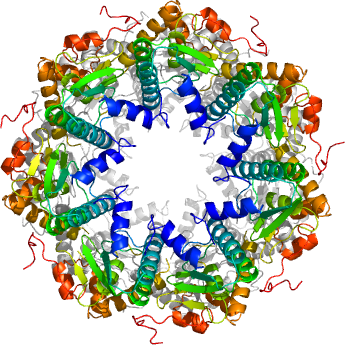
1. Smith, C., Baker, T. and Sauer, R. Lon and Clp family proteases and chaperones share homologous substrate-recognition domains. Biochemistry 96 (12), 6678-6682, (1999).
2. Brotz-Oesterhelt, H., Beyer, D., Kroll, H., Endermann, R., Ladel, C., Schroeder, W., Hinzen, B., Raddatz, S.,
Paulsen, H., Henninger, K., Bandow, J., Sahl, H. and Labischinski, H. Dysregulation of bacterial proteolytic machinery
by a new class of antibiotics. Nature Medicine, 11 (10) 1082-1087 (2005).
3. Wang, J., Hartling, J. and Flanagan, J. The Structure of ClpP at 2.3 A° Resolution Suggests a Model for ATP-Dependent Proteolysis. Cell, Vol. 91, 447–456, 1997.
4. Kanga, S., Maurizia, M., Thompson, M., Mueserb, T. and Ahvazib, B. Crystallography and mutagenesis point to an essential role for the N-terminus of human mitochondrial ClpP. Journal of Structural Biology 148, 338–352, 2004.
5. Gribun, A., Kimber, M. S., Ching, R., Spranger, R., Fiebig, K. M., & Houry, W. A. The ClpP Double-Ring Tetradecameric Protease Exhibits Plastic Ring-Ring Interactions and the N-Termini of Its Subunits Form Flexible Loops that are Essential for ClpXP and ClpAP Complex Formation. Journal of Biological Chemistry 280(16), 16185-16196 (2005).