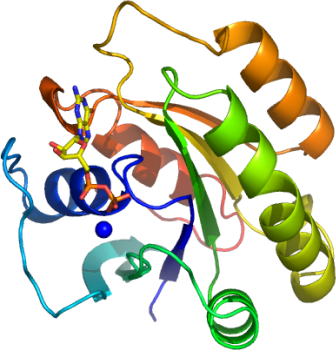
DIRAS
PDB:2GF0
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:DIRASA-s001
Entry Clone Source:MGC
SGC Clone Accession:Tag:N-terminal, TEV cleavable hexahistidine tag
Host:Bl-21(DE3)R3 phage resistant strain
Construct
Prelude:sm residues are from the construct that remain after the TEV cleavage of the hexahistidine tag.
Sequence:smPEQSNDYRVVVFGAGGVGKSSLVLRFV KGTFRDTYIPTIEDTYRQVISCDKSVCTL QITDTTGSHQFPAMQRLSISKGHAFILVF SVTSKQSLEELGPIYKLIVQIKGSVEDIP VMLVGNKCDETQREVDTREAQAVAQEWKC AFMETSAKMNYNVKELFQELLTLETRRNM SLNIDGKRSGKQKRTDRVKGKCTLM
Vector:pNIC28-Bsa4
Growth
Medium:Antibiotics:Procedure:A 20 ml starter culture (LB + 50 µg/ml kanamycin) of freshly transformed E. coli cells was used to inoculate 1 litre of TB plus 50 µg/ml kanamycin. When OD 600 reached ~1.0 the temperature was shifted down from 37°C to 25°C for 1 hour before induction with the addition of 1 mM IPTG. Protein expression was allowed to carry on for a further 4 hours before harvest.
Cells were spun at 4000 rpm for 10 mins at 4°C. The pellets resuspended in 25mls Resuspension Buffer. The resuspended cell pellets were placed in the -80°C freezer
Purification
ProcedureNi-NTA added to BioRad drip column: 4mls (50 %) Resin washed with 12.5 ml of WB1 Each supernatent was applied to a column using a 5 ml pipette. The flow through was collected in a 50 ml falcon tube and re-applied to the column. Wash with 12.5 ml of WBI and collect run through. Wash with 12.5 ml column vols of WBII and collect run through. Elute with 14 mls of EB.
At this stage the purity of the protein was greater than 90 % based on SDS - PAGE analysis. The C-terminal hexahistidine tag was removed by TEV protease treatment. The TEV protease, a hexahistidine-tagged construct, was over-expressed and purified in-house to a final concentration of 2.5 mg/ml.
Add 100 µl of the home produced TEV protease to each fraction and left at 4°C overnight. The following steps were carried out to remove the cleaved products and TEV protease: Change buffer from Elution Buffer to 50 mM Tris pH 8, 150 mM NaCl, 10 mM MgCl2 using a 10-kD cutoff concentrator. Place 200 m l of 50 % Ni-NTA agarose in a 1.5 ml eppendorf tubes, add 1ml of 50 mM Tris pH 8, 150 mM NaCl mix, spin down and remove buffer. Repeat this resin wash step once. Add the TEV treated protein sample to the resin and mix for 30 min. Finally spin down resin and collect the supernatant which contains the cleaved DIRASA.
Concentration: The concentration of DIRASA was measured and 5 molar equivalents of GTP were added before concentrating to a 24 mg/ml. The concentrated protein was aliquoted into 50 µl volumes before freezing in the -80degC freezer.
Extraction
Procedure1 proteinase inhibitor tablet (EDTA free) was added to 10ml Resuspension Buffer. This was used to syspend the pellet before homogenisation. Total vol: 45 ml (estimate).
5 passes through the Emulsiflex C5 high pressure homogeniser. Total vol: 50 mls (estimate). Centrifuge for 40 mins at 16000 rpm and 4°C to remove cell debris.Discard pellet.
Concentration:LigandMassSpec: Before His-Tag remove After His-Tag remove NAME (construct) EXPECTED MWt MEASURED MWt EXPECTED MWt MEASURED MWt DIRAS2A-c005 24881.2 24881.1 (main) 22415.6 22415.7 (main) 24832.3 22171.5 24941.7
Unresolved contaminants still present after TEV cleavage.
Crystallization:Small crystals grew from a 1:1 ratio mix of DIRASA(Mg + GTP)-to-reservoir (3.5 M sodium formate pH 7.0). Crystal size was improved by using crushed original crystals as seeds in a crystallisation setup that used the same original conditions.
NMR Spectroscopy:
Data Collection:Resolution: 1.9Å, X-ray source: Synchrotron SLS -X10, single wavelength.
Data Processing: