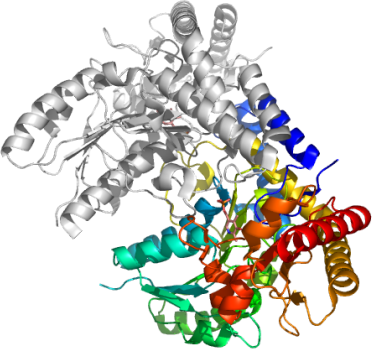
SCLY
PDB:2HDY
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:AAH07891
Entry Clone Source:
SGC Clone Accession:
Tag:
Host:
Construct
Prelude:
Sequence:SMGRDAPAPAASQPSGCGKHNSPERKVYMDYNATTPLEPEVIQAMTKAMWEAWGNPSSPYSAGRKAKDIINAARESLAKMIGGKPQDIIFTSGGTESNNLVIHSVVKHFHANQTSKGHTGGHHSPVKGAKPHFITSSVEHDSIRLPLEHLVEEQVAAVTFVPVSKVSGQTEVDDILAAVRPTTRLVTIMLANNETGIVMPVPEISQRIKALNQERVAAGLPPILVHTDAAQALGKQRVDVEDLGVDFLTIVGHKFYGPRIGALYIRGLGEFTPLYPMLFGGGQERNFRPGTENTPMIAGLGKAAELVTQNCEAYEAHMRDVRDYLEERLEAEFGQKRIHLNSQFPGTQRLPNTCNFSIRGPRLQGHVVLAQCRVLMASVGAACHSDHGDQPSPVLLSYGVPFDVARNALRLSVGRSTTRAEVDLVVQDLKQAVAQLEDQA
Vector:pNIC-Bsa4
Growth
Medium:
Antibiotics:
Procedure:30 microL competent BL-21 (DE3) cells were transformed with 2 µl plasmid miniprep for 30 min on ice followed by heatshock at 42°C for 45 sec. SOC, 125 µl, was added to the cellsuspension which was then incubated for 1 hour at 37°C and plated on LB-plates containing kanamycin (50 µg/mL). 20 mL TB with 100 µg kanamycin/mL was inoculated with cells and grown overnight (ON) at 30°C. The inoculation culture was added to 1.5 L TB (supplemented with 50 µg kanamycin/mL) in 2 L bottles. The flask was incubated in the LEX system-water bath at 37°C until OD600 reached 1. At this time the flask was transferred to an 18°C water bath in the LEX-system. Expression of protein was induced by addition of 0.5 mM IPTG and continued for approximately 18 hours.
Cells were harvested by centrifugation in a SLC-6000 rotor for 10 minutes at 5000 rpm (OD600 9.1; WCW 23.3 g). Pellets were suspended in 100 ml 50 mM Nafosfate pH 7.5, 10 % glycerole, 0.5 mM TCEP and 500 mM NaCl, 10mM imidazole and Complete EDTA-free protease inhibitor (Roche Biosciences). Suspended cells were stored at -80°C until further use. Before lysis, 4 µl of 250U/µl benzonase (Novagen) was added to the suspended cells.
Purification
Procedure
SCLYA was purified on a Hi-Trap chelating column followed by a Superdex 200 gel filtration column on the Äkta Express. The protein migrated as a well defined single peak and fractions containing protein were coloured bright yellow from the bound PLP co-factor. These fractions were pooled and the TCEP-concentration adjusted to 2 mM before concentration in AmiconUltra filters to 22.8 mg/mL. The protein was aliquoted and quick-frozen in liquid nitrogen.
TEV-cleavage: TEV-cleavage was performed by adding 422 µl His-tagged TEV (30 µM) to 30 mg protein in a total volume of 1.8 ml in gel filtration buffer with 2 mM TCEP for approximately 72 hours at 4°C. The sample was diluted with 20 mM Hepes, 500 mM NaCl, 10% glycerole and 0.5 mM TCEP at pH 7.5 before loading it onto a His-trap crude column on an Äkta Prime. The cleaved protein bound weakly to the column and was eluted with the same buffer plus 35 mM imidazole. After elution the TCEP concentration was adjusted to 2 mM and the sample concentrated to 35.6 mg/ml in a AmiconUltra MWCO 30000 concentrator.
Extraction
Procedure
The sample was sonicated (Sonics VibraCell) at 80% amplitude for 3 min (effective time, pulse: 4Â on 4Â off).The sample was spun for 30 min at 20500 rpm in a Sorvall SA-800 rotor and the soluble fraction decanted and filtered through a 0.45 µm syringe filter
Concentration:
Ligand
MassSpec:
Crystallization:Sitting drops containing 0.1 µl of protein solution (17 mg/mL) + 0.1µl well solution containing 100 mM HEPES pH 6.7 and 10% PEG6000 was left to equilibrate against well solution. Crystals appeared after 3 days at 20 degC.
NMR Spectroscopy:
Data Collection:Data was collected at the ESRF, beam-line ID23-1. Data was processed with XDS and XSCALE. The space group is P212121 with cell parameters 59 86 189 90 90 90.
Data Processing:The structure was solved by Molecular Replacement using a monomer of the E. coli cysteine desulfurase IscS (pdb entry: 1P3W) as the search model. The asymmetric unit contains one protein dimer. Refmac was used for refinement and Coot for model building. TLS refinement with 4 TLS groups was used in Refmac. The final model starts at glutamate 29 and ends at glutamine 444, the penultimate residue in the sequence. Residues 120 to 132 in both monomers are disordered and invisible in the electron density.