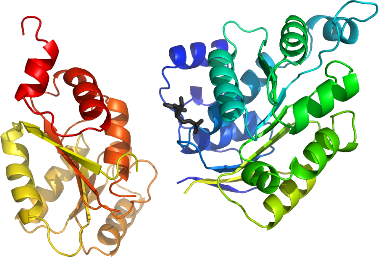
DDX3X
PDB:2I4I
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:Entry Clone Source:SGC Clone Accession:Tag:Host:Construct
Prelude:Sequence:SMVEATGNNCPPHIESFSDVEMGEIIMGNIELTRYTRPTPVQKHAIPIIKEKRDLMACAQTGSGKTAAFLLPILSQIYSDGPGEALRAMKENGRYGRRKQYPISLVLAPTRELAVQIYEEARKFSYRSRVRPCVVYGGADIGQQIRDLERGCHLLVATPGRLVDMMERGKIGLDFCKYLVLDEADRMLDMGFEPQIRRIVEQDTMPPKGVRHTMMFSATFPKEIQMLARDFLDEYIFLAVGRVGSTSENITQKVVWVEESDKRSFLLDLLNATGKDSLTLVFVETKKGADSLEDFLYHEGYACTSIHGDRSQRDREEALHQFRSGKSPILVATAVAARGLDISNVKHVINFDLPSDIEEYVHRIGRTGRVGNLGLATSFFNERNINITKDLLDLLVEAKQEVPSWLENMAYEHHYKG
Vector:pNIC-Bsa4
Growth
Medium:Antibiotics:Procedure:30 µl competent BL-21 (DE3) cells were transformed with 2 µl plasmid miniprep for 30 min on ice followed by heatshock at 42°C for 45 sec. SOC, 125 µl, was added to the cellsuspension which was then incubated for 1 hour at 37°C and plated on LB-plates containing kanamycin (50 µg/mL). 20 mL TB with 100 µg kanamycin/mL was inoculated with cells and grown overnight (ON) at 30°C. The inoculation culture was added to 1.5 L TB (supplemented with 50 µg kanamycin/mL) in 2 L bottles. The flask was incubated in the LEX system-water bath at 37°C until OD600 reached 1. At this time the flask was transferred to an 18°C water bath in the LEX-system. Expression of protein was induced by addition of 0.5 mM IPTG and continued for approximately 18 hours.
Cells were harvested by centrifugation in a SLC-6000 rotor for 10 minutes at 5000 rpm. Pellets were suspended in 50 mM Nafosfate pH 7.5, 10 % glycerole, 0.5 mM TCEP, 500 mM NaCl, 10mM imidazole and Complete EDTA-free protease inhibitor (Roche Biosciences) was added (1 tablet per 100 ml, final volume 116 ml). Suspended cells were stored at -80°C until further use.
Purification
ProcedureThe cell lysate was loaded onto a HisTrap FF crude column with a peristaltic pump and purified on an Äkta purifier system. The column was washed with IMAC-wash buffer before bound protein was eluted with 500 mM imidazole. Affinity-purification was followed by gel-filtration on a Superdex 200 16/60 gel filtration column in gel-filtration buffer. DDX3X migrated as a nice single peak corresponding to a monomer form of the protein. Fractions containing protein were pooled, the TCEP-concentration adjusted to 2 mM, and concentrated to 2.44 mg/ml in AmiconUltra MWCO 30000 concentrators. The correct mass of the DDX3X construct was verified by mass spectrometry.
TEV-cleavage: The N-terminal His-tag was removed by incubating 20 mg of DDX3X with a His-tagged TEV-protease at a molar ratio of 1:50 in a final volume of 10 ml overnight at 4? C. The reaction mixture was loaded onto a HisTrap FF crude column and cleaved DDX3X was detected in the flow-through and concentrated to 23.8 mg/ml before setting up crystallization drops. SDS-PAGE analysis showed that the purified protein was more than 95% pure (results not shown).
Extraction
ProcedureBefore lysis, 8 µl of 250U/µl benzonase (Novagen) was added and lysis was performed by sonication (3Â 4ÂÂ on/4ÂÂ off, amplitude 70%). Cell debris was pelleted by centrifugation (30 min at 20500 rpm in a Sorvall SA-800 rotor). The soluble fraction was filtered through a 0.45 µm syringe filter.
Concentration:LigandMassSpec:Crystallization:DDX3X crystals were grown using vapor diffusion at 4 °C by mixing equal amounts of protein solution at a concentration of 21 mg/ml containing 20mM ATP?S or 20 mM ADP and 20mM MgCl2 and reservoir solution containing 1.75M Sodium Formate and 100mM Tris-HCl pH 7.5. Crystals appeared after one day and had reached their maximal size in twelve days (approximately 300 µm × 100 µm × 50 µm).
NMR Spectroscopy:Data Collection:Data Processing:The structure was solved by Molecular Replacement using the two domains separately. As search model for the DEAD domain, the N-terminal RecA-like domain of DjVLGB (PDB entry: 1WRB) was used and similarly for the helicase domain the ATP-dependent RNA helicase vasa (PDB entry: 2DB3) was used. The asymmetric unit contains one protein monomer. Refmac was used for refinement and Coot for model building. TLS refinement using 4 TLS groups as suggested by tlsmd (
http://skuld.bmsc.washington.edu/~tlsmd/) was used with Refmac. In the refinement, data in the interval 15-2.20Å resolution was used and the progress of refinement was followed by decreasing R and Rfree values. At the end of the refinement the values for R=0.1887 and Rfree=0.2182. The final model starts at methionine 167 and ends at tyrosine 580. Residues 407-410, 535-536 and 581-582 are disordered and not visible in the electron density. A Ramachandran plot shows 98% of all residues in favored regions and no outliers in the disallowed regions. Coordinates for the crystal structure were deposited in the Protein Data Bank, accession code 2I4I.