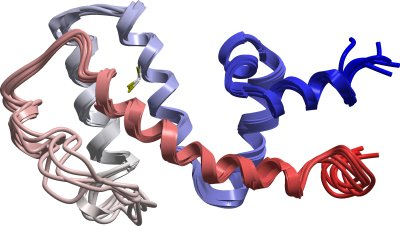
RGS10 (NMR)
PDB:2I59
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:IMAGE: 4301597
Entry Clone Source:MGC
SGC Clone Accession:Tag:N-terminal, TEV cleavable (*) hexahistidine tag. Tag sequence: MHHHHHHSSGVDLGTENLYFQ(*)SM
Host:E. coli BL21(DE3)-Rosetta
Construct
Prelude:Sequence: SMSMQSLKSTAKWAASLENLLEDPEGVKR FREFLKKEFSEENVLFWLACEDFKKMQDK TQMQEKAKEIYMTFLSSKASSQVNVEGQS RLNEKILEEPHPLMFQKLQDQIFNLMKYD SYSRFLKSDLFLKHKRTEEEEEDL
Vector:pLIC- SGC1.
Growth
Medium:Antibiotics:Procedure:BL21(DE3)-Rosetta competent cells were transformed with the expression plasmid and plated on LB plates containing 60 µg/ml carbenicillin and 30µg/ml chloramphenicol. 4 Colonies from the transformation were used to inoculate 4 times 2 ml LB containing 60 µg/ml carbenicillin and 30µg/ml chloramphenicol. The cells were then grown at 700 RPM , 37°C for 6 hrs. The temperature was reduced to 22°C and the cells were induced with 1 mM IPTG. Expression of the 4 clones was analyzed by SDS - PAGE . The best clone was used for large scale expression in either [15N]-M9 or [13C, 15N]-M9 medium, containing 0.5g [15N]-NH4Cl per L and, for 13C-labelling, 2g [13C]-glucose per L.
Cell growth and induction: Starter overnight cultures of 80 ml were grown at 37°C in either [15N]-M9 or [13C,15N]-M9 supplemented with 60 µg/ml carbenicillin and 30µg/ml chloramphenicol. The large scale cultures were grown in 4 x 450 ml M9 with 60 µg/ml carbenicillin and 30µg/ml chloramphenicol in 4 x 2 L bottles (dilution for inoculation 1: 25). The culture was grown at 37°C and transferred to 22°C when the OD600 reached a value of 0.5. The culture was induced with 1 mM IPTG and grown at 22 °C overnight. The next day the cells were harvested by centrifugation, washed with ice-cold 150 mM NaCl and frozen at -80°C.
Purification
ProcedureColumn 1 : Ni-affinity, MC-POROS, 8 ml (Applied Biosystems)
Procedure: The cell extract was loaded on the column at 1 ml/min on a workstation Vision (Applied Biosystems). The column was washed with 10 column volumes of start buffer and eluted with a gradient from 5 to 500 mM imidazole at a flow rate of 5 ml/min. The extinction at 280nm was monitored and fractions were collected and analyzed by SDS - PAGE . Positive fractions were pooled and filled into a dialysis bag with 8 kDa MW cutoff.
TEV cleavage and dialysis: The His-tag was cleaved with 1 mg TEV per 40 mg target protein in a dialysis bag and dialysed with 5 L of: 20 mM Tris HCl 8.0, 500 mM NaCl, 1 mM ß mercaptoethanol, at 15°C overnight.
Column 2 : As column 1 but the flow rate was 0.5 ml/min and without imidazole in the start buffer.
Procedure: The flow through was collected and concentrated.
Concentration and buffer exchange: Using Amicon Ultra-15 concentrators with 5 kDa cutoff, all RGS 10A-c009 samples were exchanged into NMR buffer and concentrated as follows:
- U-[15N]-labelled RGS 10A: final 1.9 mM protein in 20 mM phosphate buffer pH 7.0, 50 mM NaCl.
- U-[13C,15N]-labelled RGS 10A: final 1 mM protein in 20 mM phosphate buffer pH 6.0, 50 mM NaCl, 1 mM dDTT, 0.02% sodium azide.
Samples in D2O were prepared by lyophilization and redissolving into 100% D2O.
Concentrations were determined from the absorbance at 280 nm.
Extraction
ProcedureConcentration:LigandMassSpec:Calculated mass of the construct was 16487 Da (15N: 16676 Da /13C,15N: 17412 Da). The determined mass for the U-[15N] sample was 16638 Da and for the U-[15N,13C] sample 17364 Da.
Crystallization:NMR Spectroscopy:Data Collection: NMR spectra were acquired at 297 K, using Bruker DRX 600 and DMX 750 spectrometers in standard configuration with triple resonance probes equipped with self-shielded triple axis gradient coils. Spectra for the resonance and NOE assignment were recorded essentially as described in the original references. A 1.9 mM 15N-labelled RGS 10 sample in 90% H2O/10% D2O (NMR buffer; pH 6.0) was used for 3D 15N-separated NOESY-HSQC, 15N T1 and 15N T2 relaxation, and heteronuclear 15N-1H NOE experiments. A 1 mM 13C, 15N-labelled sample of RGS 10 in 90% H2O/10% D2O (NMR buffer; pH 6.0) was used for all HN-detected triple resonance experiments, 3D CBCA(CO)NNH, CBCANNH, CC(CO)NNH, H(CCCO)NNH, HBHA(CBCACO)NNH, HNCO, HN(CA)CO, and for 3D 13C-separated aliphatic-centred- and aromatic-centred NOESY-HSQC spectra . The sample was then freeze-dried and redissolved in 100% D2O for acquisition of 3D 13C-separated HMQC-NOESY, HCCH-COSY, HCCH-TOCSY and 2D NOESY and TOCSY spectra. Data were processed on Silicon Graphics O2 workstations using the program XWIN-NMR (version 2.6) of Bruker BioSpin GmbH (Rheinstetten , Germany ).
Assignment: Assignment of 13C, 15N and 1H resonances was carried out using standard assignment procedures on Linux workstations Intel Dual Xeon 3GHz PC, with the interactive program CCPNMR Analysis version 1.0.9 ( Vranken et al., Proteins 59, 687-696; http://www.ccpn.ac.uk/ ). The assignments are deposited in the BioMagResBank (http://www.bmrb.wisc.edu/) under accession code BMRB- 7272.

1H-15N HSQC of RGS 10 at 750 MHz and 297 K |
Data Processing:Preliminary three dimensional structures of RGS 10A were calculated using the program CYANA v. 2.0 ( Güntert et al., (1997) J. Mol. Biol. 273 , 283-298; Herrmann et al., (2002). J. Mol. Biol. 319 , 209-227 ) based on the resonance assignments, and NOE peak lists from 13C HMQC-NOESY, 3D 13C-aliphatic-centred NOESY-HSQC , 3D 13C-aromatic-centred NOESY-HSQC, and 3D 15N NOESY-HSQC spectra. Dihedral angle restraints were predicted using the program TALOS (Cornilescu et al., J. Biomol. NMR, 13 (1999) 289-302;
http://spin.niddk.nih.gov/NMRPipe/talos/) and used to aid initial rounds of structure calculations but were excluded in the final rounds. A precise ensemble of structures showing an obvious RGS fold was obtained to which hydrogen bond restraints were added on the basis of the observed protected amides following sample exchange into D2O. Iterative structure refinement was carried out manually using XPLOR-NIH v. 2.14 ( Schwieters et al., (2003) J. Magn. Reson. 160 , 66-74;
http://nmr.cit.nih.gov/xplor-nih/). The refined ensemble of the 5 lowest energy structures with no NOE violations was submitted to the PDB under code 2I59. Also included in the deposition are chemical shift list and list of NMR restraints.