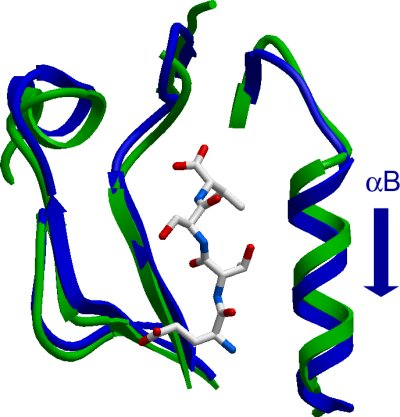
MPDZ (3rd): New alpha helix displacement: Human multiple PDZ domain protein (3rd domain)
PDB:2IWN
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:NM_003829 Variant; GenBank GI number: gi|4505231
Entry Clone Source:Origene
SGC Clone Accession:MPDZA-c113
Tag:Host:E.coli BL-21(DE3)R3 phage resistant, Rosetta strain
Construct
Prelude:Residues in lower case are the histidine tag followed by the TEV recognition sequence with the cleavage site marked with a *. ETSV = C-terminal PDZ recognition motif.
Sequence:mhhhhhhssgvdlgtenlyfq*smSETFD VELTKNVQGLGITIAGYIGDKKLEPSGIF VKSITKSSAVEHDGRIQIGDQIIAVDGTN LQGFTNQQAVEVLRHTGQTVLLTLMRRGE TSV
Vector:pNIC28-Bsa4
Growth
Medium:Antibiotics:Procedure:2 microL of construct DNA was added to a well of a PCR plate on ice. 90 microL of competent BL21(DE3)-R3/Rosetta bacteria were added, with the plate remaining on ice. The plate was left on ice for a further 20 minutes. The plate was transferred to a 42degC water bath for 45 seconds and then returned to sit on ice for 2 minutes. 100 microL of SOC medium (pre-warmed to 42degC) was added and the plate incubated at 37degC for 60 minutes.180 microL of the culture was plated out onto LB-agar containing 50 microG/mL kanamycin and 34 microG/mL chloramphenicol in a 5.5cm Petri dish. The plates were incubated at 37degC overnight. 40 mL of LB containing 50µg/ml Kanamycin and 34 microG/mL chloramphenicol was prepared in 250 mL flasks were prepared. A number of colonies, from the transformation plate, were added to each starter culture. The flask was left in a shaker at 200 rpm and 37degC overnight.18 mL of the starter culture was added to autoclaved 1litre TB medium that contained 50 microG/mL kanamycin. The flasks were placed in a shaker at 180 rpm and 37degC. After 4 hours of growth the temperature was reduced to 20degC. One hour later when the cell density reached an OD600 of approximately 1.0 the induction was initiated with the addition of 0.1 mM IPTG.The cells were spun down (4500 rpm, 15 mins). The pellets were resuspended in 30 mL lysis/ binding buffer, transfered into 50 mL tubes and placed in a -80degC freezer.
Purification
ProcedureTotal volume of Ni-NTA added to BioRad drip column: 2 mls (50%)
Resin washed with 20 ml of Binding Buffer.
The supernatent was applied to a column using 5 ml pipette and allowed to pass over the resin. The flow through was collected in a 50 ml falcon tube and applied once more to the column. The loaded resin was washed with 15 ml of Wash Buffer. Elute with ~11 mL of Elute Buffer into 6 x 2 mL fractions. 100 microL of in-house purified TEV protease was added to the pooled fractions and incubated overnight at 4degC. The following day an additional 100 microL was added due to incomplete cleavage as judged using SDS-PAGE analysis.
Enzymatic treatment : Affinity chromatography in a batch mode was applied to remove the undigested products, the cleaved histidine tags and the histadine-tagged TEV protease.
200 microL Ni-sepharose (new) was washed three times before adding to the digested protein. The slurry was rotated for 45 min at 4degC. To separate the gel from from the sup: 1. Centrifugation 5 min/500xg; 2. The supernatant was filtered through a 1.2 micrometer syringe filter.
The final concentration was done using a Vivaspin (20ml mwco 5 KD) concentrator with the protein being concentrated to 10 mg/ml before freezing and storing in a -80degC freezer.
Extraction
ProcedureAfter thawing, the resuspended cells were lysed after three passes through the Emusiflex C5 high pressure homogeniser. PEI (stock 5 %) was added to the homogenate to a final concentration of 0.15 %. The cell debris, nuclei and DNA were spun down at 16,500 rpm for 45 min.
Concentration:LigandMassSpec:After His-Tag removal: Expected MWt. = 10459.8; Measured MWt. = 10460.4
Crystallization:Crystals grew from a 1:1 ratio mix MPDZA-to-reservoir (30% PEG 10K; 0.20M Li2SO4; 0.1M acetate pH 4.5).
NMR Spectroscopy:Data Collection:Resolution: 1.35Å;X-ray source: Synchrontron SLS-X10.
Data Processing: