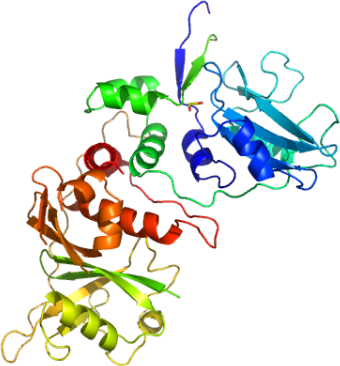
SOCS4-ElonginC-ElonginB complex
PDB:2IZV
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:gi|40807480
Entry Clone Source:
SGC Clone Accession:
Tag:N-terminal his6 tag, TEV-protease cleavable (*) however tag cannot be cleaved due to steric hindrance
Host:BL-21(DE3)R3 phage resistant
Construct
Prelude:
Sequence:SOCS4: mhhhhhhssgvdlgtenlyfqs*MLVPDLLQINNNPCYWGVMDKYAAEALLEGKPEGTFLLRDSAQEDYLFSVSFRRYSRSLHARIEQWNHNFSFDAHDPCVFHSPDITGLLEHYKDPSACMFFEPLLSTPLIRTFPFSLQHICRTVICNCTTYDGIDALPIPSSMKLYLKEYHYKSKVRVLRIDAPE
ElonginC:
MMYVKLISSDGHEFIVKREHALTSGTIKAMLSGPGQFAENETNEVNFREIPSHVLSKVCMYFTYKVRYTNSSTEIPEFPIAPEIALELLMAANFLDC
ElonginB:
MDVFLMIRRHKTTIFTDAKESSTVFELKRIVEGILKRPPDEQRLYKDDQLLDDGKTLGECGFTSQTARPQAPATVGLAFRADDTFEALCIEPFSSPPELPDVMKPQDSGSSANEQAVQ
Vector:pNIC28-Bsa4
Growth
Medium:
Antibiotics:
Procedure:A 50 ml overnight culture in LB was grown and used to inoculate 6L TB at 1:300. Plasmids were maintained by kanamycin and chloramphenicol. Cultures were grown at 37 degC until they reached an OD 600 of 0.5 and then cooled to 20 degC. Expression was induced overnight for ~14 hours using 0.5 mM IPTG at an induction OD 600 of 1.0. The cells were harvested by centrifugation, transferred to 50 ml tubes and frozen.
Purification
Procedure
Column 1: Ion exchange - Nucleic acid removal. DEAE cellulose (DE52, Whatmann), 50 g of resin divided in two 2.5 x 20 cm columns. The resin was hydrated in 2.5M NaCl, then washed with 20 ml binding buffer prior to loading the sample.
Procedure: The column was equilibrated with binding buffer. The lysate was applied to the column which was subsequently washed with wash buffer 1. CSNK1G3 was eluted with elution buffer. The eluted protein was collected and analyzed by SDS-PAGE. DTT was added to the protein sample to a final concentration of 5mM. The N-terminal his 6 -tag was cleaved by incubating the protein overnight with TEV protease, shrimp alkaline phosphatase and lambda phosphatase.
Column 2: Ni-affinity
Ni-sepharose (Amersham), 8 ml of 50% slurry in 1.5 x 10 cm column, washed with binding buffer.
Procedure: The flowthrough from column 1 was loaded by gravity flow on the Ni-sepharose column. The column was then washed with 200 ml wash buffer under gravity flow. The protein was eluted by gravity flow by applying 5-ml portions of elution buffer with increasing concentration of imidazole (50 mM, 100 mM, 150 and 250 mM); fractions were collected until essentially all protein was eluted. After elution DTT was added to a final concentration of 10 mM.
Column 3: HiLoad 16/60 S200 Gel Filtration
Procedure: Protein was applied in 25 mM HEPES pH 7.5, 250 mM NaCl and the ternary complex fractions pooled.
Column 4: Ion exchange Source 15Q column
Procedure: SOCS4-ElonginC/B complex was applied to a 10 ml Source15Q column in buffer A and eluted from the column by a linear gradient with buffer B . Elution occurred at ~250 mM NaCl.
Extraction
Procedure
The frozen cells were thawed on ice and binding buffer (plus 1 mM PMSF, 0.5 mM TCEP) added to a final volume of 350 ml. Cells were lysed using a high pressure cell disruptor. The lysate was centrifuged at 17,000 RPM for 30 minutes and the supernatant collected for purification.
Concentration:Samples containing the ternary complex were pooled and concentrated to 11 mg/ml using Centricons (10 kDa cut off). The final buffer contained 50 mM HEPES pH 7.5, 250 mM NaCl, 10 mM DTT. Protein composition was confirmed using LC-ESI MS-Tof.
Ligand
MassSpec:Masses of purified proteins were confirmed by LC-MS, using an Agilent LC/MSD TOF system with reversed-phase HPLC coupled to electrospray ionisation and an orthogonal time-of-flight mass analyser. Proteins were desalted prior to mass spectrometry by rapid elution off a C3 column with a gradient of 5-95% acetonitrile in water with 0.1% formic acid.
Crystallization:Crystals were obtained at 4 degC in 150 nl sitting drops set up at a ratio of 1:1 with mother liquor and 11 mg/ml protein. The mother liquor consisted of 2M NaCl, 10% PEG6000.
NMR Spectroscopy:
Data Collection:
Data Processing:Data were collected at the SLS beamline 10. The structure was solved by molecular replacement using the SOCS2 complex determined previously at the SGC (pdbcode 2C9W).