5′-Nucleotidases catalyze the dephosphorylation of ribo- and deoxyribonucleoside monophosphates into their corresponding nucleoside forms (Hunsucker et al. 2005). They regulate cellular nucleotide and nucleoside levels by counteracting nucleoside kinases in substrate cycles. They are also likely to affect the phosphorylation level of nucleoside analogs used in treatment of, for example, cancer and HIV. Since most nucleoside analogs need to be phosphorylated to exert their pharmacological activity, the actions of 5′-nucleotidases are likely to reduce the pharmacological effect of these drugs.
The human cytosolic 5′-nucleotidase II (NT5C2 or cN-II) is specific for the 6-hydroxypurine nucleoside monophosphates IMP, dIMP, GMP, dGMP and XMP. It has also been called IMP-GMP specific nucleotidase and high Km 5′-nucleotidase. It possesses phosphotransferase activity, thus it is also able to transfer a phosphate to a nucleoside acceptor to form a nucleoside monophosphate (Worku & Newby, 1982). It is allosterically activated by nucleotides (preferably ATP) and 2,3-bisphosphoglycerate (2,3-BPG) and inhibited by inorganic phosphate (Spychala et al., 1988). It is a 60 kDa enzyme that functions as a tetramer and is dependent of Mg2+ for activity.
NT5C2 can dephosphorylate the anti-HIV and anti-cancer nucleoside analogs FdUMP, AZTMP, CdAMP, araGMP, BVdUMP and ddCMP (Mazzon et al., 2003). It can also phosphorylate the nucleoside analogs dideoxyinosine, tiazofuran, acyclovir and ribavirin, (Johnson & Fridland, 1989; Wu JZ et al. 2005).
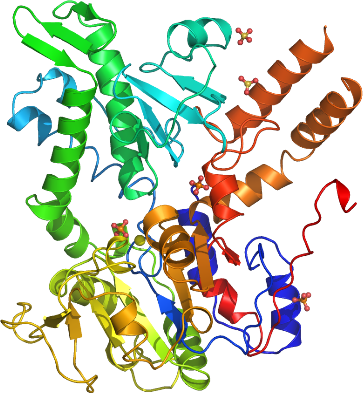
We have determined the structure of NT5C2 to a resolution of 2.2 Å. A DALI search reveals that it has similar structure to HAD superfamily enzymes, including the mitochondrial deoxyribonucleotidase, cytosolic 5’-nucleotidase III and phosphoserine phosphatase. A phosphate and a magnesium binds in the active site. The part of the active site that binds the phosphate moiety is conserved relative to other HAD superfamily enzymes suggesting similar catalytic mechanism. Two additional phosphate ions and three sulfate ions has been modeled in the structure, suggesting the location of possible effector sites.