Inosine monophosphate (IMP), the precursor of adenosine and guanosine monophosphates (AMP and GMP),
is an essential metabolite in the biosynthesis of purines.
IMP is generated predominantly by de novo biosynthesis and by interconversion of nucleotide monophosphates.
ITP is present at low levels in the cell as byproducts generated either by deamination of purine bases
or by phosphorylation of IMP.
IMP is also generated by enzymatic action of the enzyme inosine triphosphatase (ITPase; ITPA; EC 3.6.1.19)
that catalyzes the conversion of inosine triphosphate (ITP) to inosine monophosphate (IMP) and pyrophosphate (PPi).
The enzyme also catalyses the hydrolysis of dITP and XTP.
ITP --> IMP + PPi
ITPase activity prevents the cytosolic accumulation of ITP, dITP and XTP,
which may be incorporated into RNA and DNA by the respective polymerases, posing the risk of direct mutagenesis.
ITPase exists as a dimer under physiological conditions. Several naturally occurring mutations
in the ITPA gene have been found that cause ITPase deficiency.
The most common mutation is proline 32 to threonine (P32T).
Interestingly, ITPase activity is completely lost in patients homozygous for the mutation,
but reduced to 25% in patients heterozygous for the mutation.
This indicates that ITPase activity depends on the integrity of both protomers of the dimer,
and suggests that dimer activity might be regulated by crosstalk between the two protomers.
Several studies conclude that ITPase deficiency is responsible for adverse drug reactions
in patients treated with the purine analog azathioprine,
although other studies did not support such a correlation.
Azathioprine is an immunosuppressive drug used in the treatment of inflammatory bowel diseases,
leukemia, and autoimmune complications in connection with organ transplants.
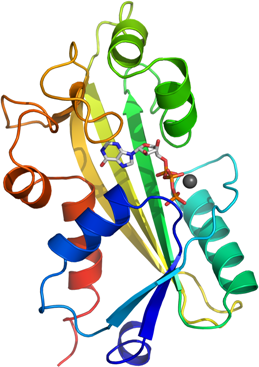
We have solved the structure of human ITPA without any ligand bound to 1.1Å resolution.
We have also solved the substrate complex to 2.8 Å.
We here identify the metal binding site and substrate binding site of this protein family.
In the ITP complex structure (PDB entry 2J4E), one molecule of ITP is bound in the cleft
between the dimerization lobe and N-terminal lobe of the protein.
The purine base is wedged between the Phe149 and Trp151 side chains of the dimerization lobe.
The substrate binds by an induced fit mechanism,
the active site cleft is closed by a large movement of an N-terminal helix. This positions several conserved residues for coordination of the substrate. The ITPases are dependent on Mg2+ or Mn2+ for activity, the location of this binding site has previously not been known. In our ITP complex we see this metal bound between Glu44 and the triphosphate tail of ITP.
<?
if (strtolower($target)=="2car") print "
Apo structure";
if (strtolower($target)=="2j4e") print "
Binary structure with ITP";
?>
Superposition of the apo (grey) and ITP-bound (pink) structures.
The upper dimerization domain and the lower N-terminal domain form a cleft that closes upon substrate binding.
ITP is shown as sticks
Arg 178, His177 and Lys 172 are within hydrogen bonding distance from the 6-keto oxygen of the inosine ring. These residues constitute a positively charged environment that interacts with the 6-keto oxygen. In ATP this position of the inosine ring is occupied by an amino group, this explains the high specificity for ITP over ATP. Because of the high levels of ATP in the cell it is essential that the enzyme does not break down this nucleotide.
In one of the eight protein chains in the asymmetric unit (D) the density showed that
the hydrolysis reaction had proceeded, and IMP and pyrophosphate occupied the active site.
The alpha- and beta-phosphates have moved apart, but otherwise no major structural rearrangements can be seen.
The P32T mutation causing ITPA deficiency is located in a loop region after the helix
that closes the active site upon substrate binding.
This loop constitutes a hinge region, and we believe that the P32T mutation
leads to a partial miss folding that prevent the dynamics necessary for substrate binding or substrate release.
Glu44 is located very close to the dimer interface; Glu44 and neighboring residues make hydrogen bonds
over the interface (Glu44, Tyr45, Gln46).
This interaction could transmit the structural problems in one monomer carrying the P32T mutation
to the other (wild type) monomer of the dimer.
This would explain the observation that individuals heterozygous for the mutations only retain 25% of ITPA activity;
which would indicate that the heterodimer between mutated and wild type ITPA is inactive.