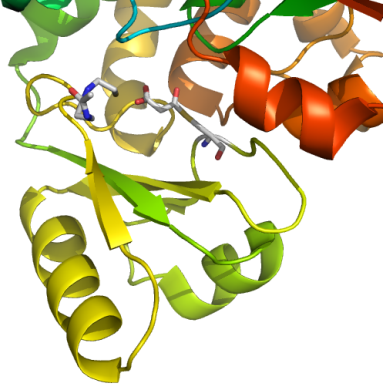
nnFAS transferase domain + CoA
PDB:2JFK
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:FASNA-s002 (BC063242)
Entry Clone Source:MGC
SGC Clone Accession:Tag:N-terminal TEV-cleavable (at *) his-tag with the following sequence mhhhhhhssgvdlgtenlyfq*s
Host:BL21(DE3)-R3
Construct
Prelude:Sequence:mhhhhhhssgvdlgtenlyfqSMRLLRASGRTPEAVQKLLEQGLRHSQDLAFLSMLNDIAAVPATAMPFRGYAVLGGERGGPEVQQVPAGERPLWFICSGMGTQWRGMGLSLMRLDRFRDSILRSDEAVKPFGLKVSQLLLSTDESTFDDIVHSFVSLTAIQIGLIDLLSCMGLRPDGIVGHSLGEVACGYADGCLSQEEAVLAAYWRGQCIKEAHLPPGAMAAVGLSWEECKQRCPPGVVPACHNSKDTVTISGPQAPVFEFVEQLRKEGVFAKEVRTGGMAFHSYFMEAIAPPLLQELKKVIREPKPRSARWLSTSIPEAQWHSSLARTSSAEYNVNNLVSPVLFQEALWHVPEHAVVLEIAPHALLQAVLKRGLKPSCTIIPLMKKDHRDNLEFFLAGIGRLHLSGIDANPNALFPPVEFPAPRGTPLIS
Vector:pNIC28-Bsa4
Growth
Medium:TB
Antibiotics:Procedure:An overnight culture (10 ml) was used to innoculate 1L TB medium (supplemented with 50 microG/ml of Kanamycin and 34 µg/ml Chloramphenicol). The cells were cultured in 2 litres at 37°C with vigorous shaking (160 rpm) until the culture reached an OD600 of 1.5. At that point temperature was reduced to 18 degC, and cells were induced with IPTG at a concentration of 0.5 mM, and cultivated further for 16 hours. Cells were harvested at 6000 rpm for 10 minutes and the cell pellet was stored at -20 degC until further use.
Purification
ProcedureColumn 1 : Ni-affinity, HisTrap, 1 ml (GE/Amersham Biosciences )
Procedure: AKTA Xpress Affinity/Gel Filtration. The cell extract was loaded on the column at 0.8 ml/minute on an AKTA-express system (GE/Amersham). The column was then washed with 10 volumes of lysis buffer, 10 volumes of wash buffer, and then eluted with elution buffer at 0.8 ml/min. The eluted peak of A280nm was automatically collected.
Column 2 : Hiload 16/60 Superdex 200 prep grade 120 ml (GE/Amersham Biosciences)
Procedure: AKTA Xpress Affinity/Gel Filtration. The eluted fractions from the Ni-affinity Histrap columns were loaded on the gel filtration column in GF buffer at 0.80 ml/min. Eluted proteins were collected in 2 ml fractions.
Concentration : The protein was concentrated to 13.7 mg/ml by using Vivaspin, 10K concentrator.
Modifications: Prior to crystallization, the protein was malonylated by addition of 10 mM malonyl CoA, 10 mM MgCl2. The mixture was incubated for 30 min at room temperature, briefly centrifuged and brought into crystallization screens. Covalent modification was verified by ESI-TOF MS.
Extraction
ProcedureThe frozen cell pellet was resuspended in 25 ml of extraction buffer. The sample was homogenised by using Emulsiflex-05 homogenizer (Glen Creston) and then centrifuged at 17000 rpm for 45 min.
Concentration:LigandMassSpec:The experimentally determined mass was 47589 Da, which corresponds with a malonyl adduct of a native protein of 47502 Da.
Crystallization:Initial screens set up with native FASN yielded several hits. However, to decrease crystal degradation by crystal handling a condition was selected that provided sufficient cryoprotection After optimisation, diffraction quality rod-like crystals were obtained from 20% PEG10K, 0.04 M Na/KPO4 and 25% glycerol with average dimensions 0.6*0.2*0.2 mm3, and were mounted with a loop and flash-frozen by plunging into liquid nitrogen. Datasets were collected at the PXII beamline at the Swiss Light Source using a MAR225 detector at 0.9793 Å. Selenomethionine-labelled protein could be crystallised using the same crystallisation condition, and crystals were analogous to the native in morphology and size but degraded after one week and had to be mounted immediately after they appeared. A SAD dataset was collected at the PXII beamline at the SLS, using the selenium peak wavelength (0.9789 Å, determined from a fluoresence scan).
NMR Spectroscopy:Data Collection:The diffraction pattern for both the native and selenomethionine-labelled protein was highly anisotropic, but images could be processed with XDS. Different scans were scaled together using XSCALE.
Data Processing:Due to severe anisotropy the anomalous signal obtained from the selenomethione-labelled crystal was significant to 4.5 Å only. However, this proved to be sufficient for SHELXD to locate 40 selenium positions. These were refined by SOLVE and solvent-flattened by RESOLVE to yield and interpretable map. NCS operators could be derived from selenium positions, but NCS averaging did not improve maps, possibly due to translational-only symmetry. Although an initial model could be built, the sequence could not be assigned and refinement stalled.
Phases obtained from this model were therefore used as external phase information in SHARP, which allowed the completion of the anomalous substructure by finding minor selenium sites. Phases were solvent-flattened and the model was rebuilt using the new map. Phase restraints (solvent-flattened phases expressed as Hendrickson-Lattman coefficients) proved very useful to keep refinement stable and improved maps significantly. After several rounds of rebuilding and refinement, the new model was used to solve the higher resolution native dataset, and was refined to convergence.