PARC
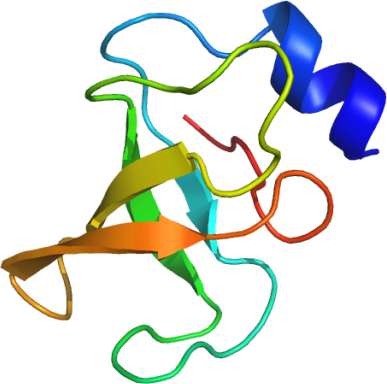
PDB:2JUF
Entry Clone Accession:NP_055904.1
Tag:mgsshhhhhhssglvpr*gshm
Host:E. coli BL21(DE3)-R3
Vector:pET 15b (Novagen)
Sequence: gshmRSEFSSRSGYGEYVQQTLQPGMRVRMLDDYEEISAGDEGEFRQSNNGIPPVQVFWQSTGRTYWVHWHMLEILGPEEATEDKASAAVEKGAGATVLGTAFPS
Growth
Procedure: The domain was expressed as a histidine (His)-fusion protein, grown in BL21(DE3) cells at 37°C for 3 hrs. The cells were induced at an OD600nm value of 0.5-1.0 using 1 mM isopropyl b-D-thiogalactopyranoside (IPTG) and incubated at 25°C for a further 5 h before harvesting. Expression was analyzed by SDS-PAGE. A uniformly 15N-labelled sample of the PARC CPH domain was prepared by growing cells in M9 defined minimal medium containing 0.8g/L 15NH4Cl and D-glucose (4g/L). To produce a 13C,15N-labelled sample, D-glucose was replaced by 13C6-D-glucose. Starter overnight cultures of 50 mL were grown at 37°C in LB medium supplemented with 100 µg/mL ampicillin. The large scale cultures were grown in 6 L Erlenmeyer flasks containing 2L of M9 defined medium supplemented with 15NH4Cl and 13C6-D-glucose (or D-glucose for uniformly 15N-labelled sample) medium with 100 µg/mL ampicillin. The culture was grown at 37°C to an OD 600nm value of 0.5-1.0. The culture was induced with 1 mM IPTG and grown at 25°C for a further 5 h. Cells were harvested by centrifugation and frozen at -20°C.
Purification
Procedure:
Column: 26/60 HiLoad Superdex 75 column (Pharmacia)
Talon (BD) was equilibrated with buffer B1. The cell lysate was adsorbed to the Talon matrix and was washed with 25 column volumes each of Buffer B1, and Buffer B2. The protein was eluted with Elution Buffer. Protein-containing fractions were analyzed by SDS-PAGE, pooled and concentrated using Centricon 10 cartridges (Amicon).
Thrombin-cleavage: The His-tag was cleaved from the fusion protein (~ 10 mg/mL) with 5 U thrombin (CalBiochem) per mg fusion protein by incubation at 4°C overnight. Following cleavage, 2 mM benzamidine was added and the cleaved tag was separated from the protein using a 26/60 HiLoad Superdex 75 column (Pharmacia) equilibrated with Gel Filtration Buffer. Gel filtration was performed at a flow rate of 2.5 mL/min and extinction at 280 nm recorded. Peak fractions were analysed by SDS PAGE, pooled and concentrated with buffer exchange using Centricon 10 cartridges (Amicon).
Extraction
Procedure: Frozen cell pellets from 2 L culture (ca. 2.5 g wet mass) were resuspended in a total volume of 50 mL ice-cold lysis buffer containing 10 mg/mL of lysozyme. Cells were then lysed by sonication.
Concentration:PARC-CPH was exchanged into NMR buffer using Amicon Centricon-10 concentrators with 10 kDa cutoff, and concentrated to 0.7mM protein. Protein sample concentrations were determined from the absorbance at 280 nm.
Structure Determination
NMR Spectroscopy: NMR spectra were acquired at 25 degC, using Bruker Avance 500 and Varian INOVA 600 spectrometers equipped with triple resonance cryoprobes and cold probe, respectively. Spectra for the resonance and NOE assignment were recorded essentially as described in the original references. A 0.7 mM 15N-labelled sample in 90% H2O/10% D2O (NMR buffer; pH 7.5) was used for 1H/15N HSQC and heteronuclear 15N-1H NOE experiments. A 0.7 mM 13C,15N-labelled sample of in 90% H2O/10% D2O (NMR buffer; pH 7.5) was used for all HN-detected triple resonance experiments, 3D CBCA(CO)NNH, CBCANNH, CC(CO)NNH, H(CCCO)NNH, HBHA(CBCACO)NNH, HNCO, HN(CA)CO, and for a 3D 13C-separated, aliphatic-centered NOESY-HSQC spectrum. The sample buffer was then exchanged to NMR buffer containing 100% D2O for acquisition of 3D 13C -NOESY, HCCH-TOCSY and TOCSY spectra. Data were processed with NMRPipe software and analyzed with the SPARKY program (Goddard, T.D., and Kneller, D.G., SPARKY 3, University of California, San Francisco). Linear prediction in the 13C and 15N dimensions was used to improve digital resolution.
Data Processing:
Assignment: Assignment of 13C,15N and 1H resonances was carried out using standard assignment procedures on Sun microsystem Ultra 5 workstations. The assignments are deposited in the BioMagResBank (http://www.bmrb.wisc.edu/) under accession code BMRB-15509.
Assigned 15N-HSQC spectrum of the human PARC-CPH domain (residues 366-466) recorded at 25 °C and 500 MHz.
Structure Calculation: Preliminary three dimensional structures were calculated using the program CYANA v. 2.0 ( Güntert et al., (1997) J. Mol. Biol. 273 , 283-298; Herrmann et al., (2002). J. Mol. Biol. 319 , 209-227 ) and were based on the resonance assignments together with NOE peak lists from 3D 13C-aliphatic-centred NOESY-HSQC , 3D 13C-aromatic-centred NOESY-HSQC, and 3D 15N NOESY-HSQC spectra. Dihedral angle restraints were derived from the chemical shifts data using the program TALOS (Cornilescu et al., J. Biomol. NMR, 13 (1999) 289-302; http://spin.niddk.nih.gov/NMRPipe/talos/). The 20 lowest-energy structures were refined using CNS by performing a short constrained molecular dynamics simulation in explicit solvent (Linge et al (2003) Proteins, 50:496-506). Structures validation was performed using MOLMOL (Koradi et al (1996) J. Mol. Graph., 14: 51-55), the Protein Structure Validation Suite (PSVS) http://www-nmr.cabm.rutgers.edu/PSVS and PROCHECK-NMR (Laskowski, R.A. (1996) J. Biomol. NMR 8:477-486). The refined ensemble of the 20 lowest energy structures with no NOE violations was deposited in the PDB under code 2juf, together with the NMR restraints used for the structure calculations.