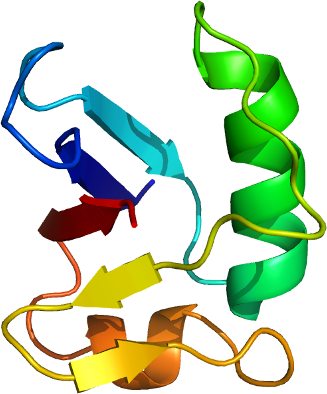
HERPUD2
PDB:2KDB
Entry Clone Accession:NP_071768.2
Entry Clone Source:ubh55.BC005091.MGC.AU35-E2.pOTB7
SGC Clone Accession:ubh55.009.085.108D07 (SDC108D07)
Tag:N-terminal: MGSSHHHHHHSSGRENLYFQ*GH
Host:BL21 (DE3)RIL
Vector:pET28MHL
Sequence: PVTLIIKAPNQKYSDQTISCFLNWTVGKLKTHLSNVYPSKPLTKDQRLVYSGRLLPDHLQLKDILRKQDEYHMVHLV
Growth
Procedure: Bacteria were grown in M9 minimal media supplemented with biotin, thiamine, and 10 uM ZnSO4; 15NH4Cl and 13C-glucose were the sole nitrogen and carbon source. Starter cultures (50 mL in a 250 mL flasks) were prepared with media supplemented with 100 uL of glycerol stock and shaken overnight (18 hours) at 220 rpm at 37 degC. They were used to inoculate 500 mL of growth media in a modified LEX fermentation system at 37 degC to an OD600 of 1.0. Cultures were induced with 1 mM IPTG and grown at room temperature overnight (15.5 hours). Cells were harvested by centrifugation and frozen in 50 mL Falcon tubes at -80 degC.
Purification
Procedure: Lysate was clarified by centrifugation for 20 min at 4 °C and the supernatant was mixed for 20 minutes at 4 °C with 2 mL settled Ni2+ affinity beads. Beads were batch-washed twice with 5 mL of cold wash buffer (spun at 2000 rpm for 6 minutes), transferred to a column, and further washed with 2 mL of wash buffer. Protein was eluted with 5 mL of elution buffer. Buffer exchange into NMR buffer and protein concentration was performed using VivaSpin concentrators with a 5,000 molecular weight cut-off at 3000 rpm, resulting in a final volume of 300 ul. Protein samples were transferred to a 5 mm shigemi NMR tube for data collection.
Extraction
Procedure: Frozen cell pellets from 500 ml cultures were thawed, resuspended in 25 mL lysis buffer, and lysed by sonication.
Structure Determination
NMR Spectroscopy:A series of spectra (13C-edited aliphatic NOESY, (13C-edited aromatic NOESY, (15N-edited NOESY-HSQC, (13C Constant Time HSQC, 3D HNCO, 3D HNCA, 3D CBCA(CO)NH, 3D HBHA(CO)NH, 3D (H)CCH-TOCSY, and 3D H(C)CH-TOCSY) were collected at 25 degC on Bruker Avance 800 MHz or 600 MHz equipped with a z-shielded gradient triple resonance cryoprobe. Chemical shifts were referenced to external DSS. All spectra were non-uniformly sampled, and were processed using the NMRPipe, NMRDraw and multidimensional decomposition software. The backbone assignments were obtained using HNCO, CBCA(CO)NH, HBHA(CO)NH, HNCA and 15N-edited NOESY-HSQC spectra. Assignments were made with the automated program FAWN. Aliphatic side chain assignments relied on H(C)CH-TOCSY and (H)CCH-TOCSY.
Distance restraints for structure calculations were derived from cross-peaks in 15N-edited NOESY-HSQC, 13C-edited aliphatic and aromatic NOESY-HSQC spectra. Automated NOE assignment and structure calculations were performed using the noeassign module implemented in CYANA. The quality of noeassign/CYANA calculation was assessed by NMR structure quality assessment scores (NMR PRF scores). The best 20 of 100 CYANA structures from final cycle were selected and subjected to molecular dynamics simulation in explicit water by the program CNS. The structures were inspected by PROCHECK and MolProbity using NESG validation software package PSVS.