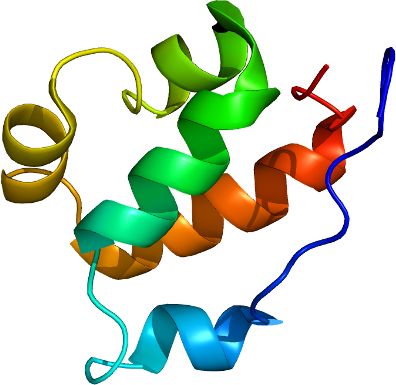
FKBP3
PDB:2KFV
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:NP_002004
Entry Clone Source:fkbp03.BC020809.MGC.AU84-G12.pDNR-LIB
SGC Clone Accession:fkbp03.001.073.031F07 (SDC031F07)
Tag:N-terminal: MGSSHHHHHHSSGLVPRGS
Host:BL-21(DE3) CodonPlus
Construct
Prelude:
Sequence:mgsshhhhhhssglvprgsMAAAVPQRAWTVEQLRSEQLPKKDIIKFLQEHGSDSFLAEHKLLGNIKNVAKTANKDHLVTAYNHLFETKRFK
Vector:pET28aLIC
Growth
Medium:
Antibiotics:
Procedure:A 250 ml flask containing M9 base minimum media (with 13C-glucose and 15N (NH4)2SO4) supplemented with 50 ug/ ml kanamycin (BioShop Canada KAN 201) was inoculated from a fresh transformed plate. The flask was shaken overnight (16 hours) at 250 rpm at 37 degC.
Using the Lex system, a 2L bottle (VWR 89000-242) containing 1800 ml of minimum media supplemented 50 ug/ ml kanamycin and 600 ul antifoam 204 (Sigma A-8311) was inoculated with 50 ml overnight LB culture, and incubated at 37 degC. The temperature of the media was reduced to 15 degC one hour prior to induction and induced at OD(600) = 2 with 100 uM isopropyl-thio-β-D-galactopyranoside (BioShop Canada IPT 001). Cultures were aerated overnight (16 hours) at 15 degC, and cell pellets collected by centrifugation and frozen at -80 degC.
Purification
Procedure
Unclarified lysate was mixed with 2-3 mL of Ni-NTA superflow Resin (Qiagen) per 40 mL lysate. The mixture was incubated with mixing for at least 45 minutes at 4 degC. The mixture was than loaded onto an empty comlum (BioRad) and washed with 100 ml wash buffer. Samples were eluted from the resin by exposure to 2-3 column volumes (approx. 10-15 mL) of elution buffer.
Gel filtration chromatography: An XK 26x65 column (GE Healthcare) packed with HighLoad Superdex 75 resin (GE Healthcare) was pre-equilibrated with gel filtration buffer for 1.5 column volumes using an AKTA explorer (GE Healthcare) at a flow rate of 2.5mL/min. The eluate sample from the IMAC step (approx. 15 mL) was loaded onto the column at 1.5 mL/min, and 2mL fractions were collected into 96-well plates (VWR 40002-012) using peak fractionation protocols). Fractions observed by a UV absorption chromatogram to contain the protein were pooled.
Extraction
Procedure
Frozen cell pellet contained in bags (Beckman 369256) obtained from 2L of culture were thawed by soaking in warm water. Each cell pellet was resuspended in 25-40 mL lysis buffer and homogenized using an Ultra-Turrax T8 homogenizer (IKA Works) at maximal setting for 30-60 seconds per pellet. Cell lysis was accomplished by sonication (Virtis408912, Virsonic) on ice: the sonication protocol was 10 sec pulse at half-maximal frequency (5.0), 10 second rest, for 10 minutes total sonication time per pellet.
Concentration:Purified proteins were concentrated using 15 mL concentrators with a 5,000 molecular weight cut-off (Amicon Ultra-15, UFC900524, Millipore) at 3750 rpm, typically resulting in a final concentration of 4-5 mg/mL.
Ligand
MassSpec:
Crystallization:
NMR Spectroscopy:
Data Collection:A series of spectra (3D 1H-13C NOESY, 3D 1H-15N NOESY, 2D 1H-13C Constant Time HSQC, 3D HNCO, 3D HNCA, 3D CBCA(CO)NH, 3D HBHA(CO)NH, 3D 1H-13C Aromatic NOESY, 3D (H)CCH-TOCSY, and 3D H(C)CH-TOCSY) were generated using a 500MHz Bruker AVANCE spectrometer, a 600MHz Bruker AVANCE spectrometer and a 800MHz Bruker AVANCE spectrometer. Spectra of aligned and unaligned spectra (2D 1H-15N IPAP HSQC) were obtained using the 500MHz Bruker AVANCE spectrometer and the 800MHz Bruker AVANCE spectrometer. NMR data was processed and analyzed using Topspin, NMRPipe, NMRDraw, Sparky, Abacus, FMC-GUI, CNS, TALOS, PALES, PSVS, and WhatIF.
Data Processing:Residual dipolar coupling sample alignment: After data collection was performed on the unaligned sample, the purified protein was aligned by titrating 12 mg/ml Pf1 co-solvent Protease-free Phage into the NMR sample until 10 Hz proton splitting was observed.