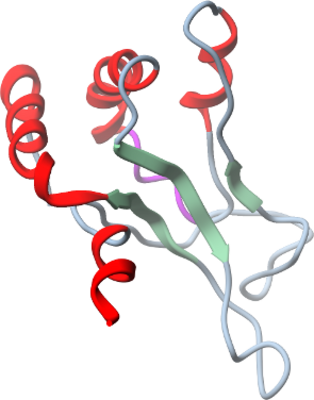
HLTF
PDB:2L1I
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:Mammalian Gene Collection cDNA template
Entry Clone Source:hltf.BC044659.MGC.AT70-E3.pCMV-SPORT6
SGC Clone Accession:hltf.52.171.179F03 (SDC179F03)
Tag:MGSSHHHHHHSSGLVPRGS
Host:E.coli BL21 CodonPlus
Construct
Prelude:
Sequence:MGSSHHHHHHSSGLVPRGSDEEVDSVLFGSLRGHVVGLRYYTGVVNNNEMVALQRDPNNPYDKNAIKVNNVNGNQVGHLKKELAGALAYIMDNKLAQIEGVVPFGANNAFTMPLHMTFWGKEENRKAVSDQLKKHGFKL
Vector:pET28a-LIC vector (GenBank, EF442785)
Growth
Medium:2 L of doubly labeled minimal media
Antibiotics:
Procedure:Competent E.coli BL21 CodonPlus cells were grown in 2 L of doubly labeled minimal media at 37 °C to OD600=1.0, then cells were induced with 1mM IPTG for 16 hrs at 15 °C.
Purification
Procedure
Cleared cell lysate in lysis buffer was loaded onto a 5 mL TALON metal-affinity resin column equilibrated in wash buffer at 4 °C temperature. The column was washed with 50 mL of wash buffer and the protein was eluted with 10 mL of elution buffer. Upon addition of 1 unit of Thrombin protease, 2 mM beta-mercaptoethanol, the eluant was incubated for 10 hrs at 4 °C. The protein was further purified by gel filtration on a HighLoad Superdex 75 column equilibrated with gel-filtration buffer. Fractions containing protein were pooled and concentrated using Amicon Ultra centrifugal filter with 3kD cutoff membrane to a final concentration of 16 mg/ml. Protein yield was 7 mg per liter of bacterial culture. Coomassie-stained SDS-PAGE showed that the 15N,13C labeled product was pure and mass-spectroscopy by LCMS (Agilent 1100 Series) showed that the purified protein had a molecular weight corresponding to 14266 g/mol.
Extraction
Procedure
Concentration:
Ligand
MassSpec:
Crystallization:
NMR Spectroscopy:NMR analyses were performed using a Bruker AVANCE 600 or 800 spectrometer at 298K. NMR samples contained 1.6 mM 15N, 13C-labeled hltf.52.171 in 20 mM sodium phosphate pH 6.3 and 100 mM NaCl with 10% D2O. The 3D spectra were measured with nonuniform sampling (NUS) of the indirect dimensions (down to 30% sampling) and reconstructed using multidimensional decomposition (MDD) using MDDNMR (Jaravine and Orekhov, 2006). All spectra were processed using NMRPipe/NMRDraw (Delaglio et al., 1995) and analyzed using Sparky (Goddard, 2008). Backbone assignments were obtained using standard triple-resonance experiments (Bax and Grzesiek, 1993). Aliphatic side-chain assignments were obtained from 3D HCCH-TOCSY, and aromatic side-chain assignments were obtained from 2D (Hβ)Cβ(CγCδ)Hδ and (Hβ)Cβ(CγCδCε)Hε experiments (Yamazaki et al., 1993). Distance restraints, derived from a 15N- and 13C-edited NOESY-HSQC (using crossrelaxation mixing time of 100 ms) and dihedral angle restraints, predicted from chemical shifts with the software TALOS (Cornilescu et al., 1999) and (Jaravine and Orekhov, 2006), were used as input for combined automated NOE assignment and structure calculation with the program CYANA (Guntert, 2004).
Data Collection:
Data Processing:The coordinates and structure factors have been deposited 2010.07.28 into the RCSB PDB database with ID code 2L1I with the following authors: Irina Bezsonova, Dante Neculai, Johan Weigelt, Chas Bountra, Aled M. Edwards, Cheryl H. Arrowsmith, and Sirano Dhe-Paganon.