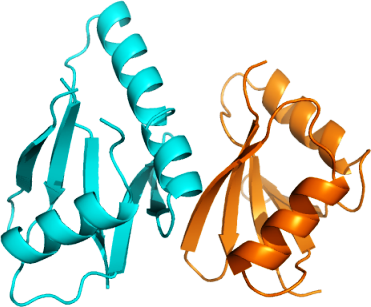
MAP2K5 + MAP3K3: Human MAP2K5 (MEK5)-MAP3K3 (MEKK3) Phox domain complex
PDB:2O2V
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:MAP2K5: NP_660143.1
MAP3K3: NP_002392
Entry Clone Source:MAP2K5: MGC
MAP3K3: MGC
SGC Clone Accession:Tag:Tag sequence: mhhhhhhssgvdlgtenlyfq^S(M); TEV-cleavable (*) N-terminal his6 tag.
Host:BL21 (DE3)
Construct
Prelude: MAP2K5A
Sequence:mhhhhhhssgvdlgtenlyfq^SMALGPF PAMENQVLVIRIKIPNSGAVDWTVHSGPQ LLFRDVLDVIGQVLPEATTTAFEYEDEDG DRITVRSDEEMKAMLSYYYSTVMEQQVNG QLIEPLQIFPRA
Vector:pNIC28-Bsa4
Growth
Medium:Antibiotics:Procedure:Grow starter cultures from freshly transformed colonies in 10 ml LB , 50 mg/ml kanamycin. This started culture was diluted 1:1000 in fresh media and was grown at 37°C to a OD600 of 0.4 and than transferred to 18°C. Expression was induced at an OD600 of 0.6 - 0.7 using 1 mM IPTG (final concentration). Cells were harvested after 4h by centrifugation (15min, 6500rpm on a JLA 8.100 rotor), transferred to 50-ml tubes, and frozen at -20°C.
Purification
Procedure Column 1 : DE52/Ni-NTA
Procedure (Gravity feed chromatography): A DE52 column (10gr suspended in 100ml of 2.5M NaCl) was equilibrated with 100ml of Loading Buffer. A 5ml NiNTA column was equilibrated with 20ml of Loading Buffer. The lysed sample was applied to the DE-52 column and washed through with 50 ml loading buffer. The flow through was applied to the 5 ml Ni-NTA column which was washed with 2x10ml of wash buffer and eluted with elution buffer in 5 ml aliquots (Step elution using 50, 100, 150, 200 and 250 mM imidazole in the Elution Buffer)
Enzymatic treatment: Treated the IMAC elution(s) with TEV protease overnight.
Column 2: SEC (AKTA-prime)
Procedure (Gel Filtration): Each component of the complex was treated separately. Fractions containing MAP3K3B or MAP2K5 collected from IMAC and treated with TEV protease overnight (identified by SDS PAGE) were concentrated to about 1ml and directly applied to a S75 16/60 column equilibrated in 10 mM Hepes pH 7.5, 100 m NaCl. The flow rate was 1ml/min and the pure proteins eluted at 72-87min (MAP3K3) and 70-85min (MAP2K5).
Concentration: The combined samples from the SEC column (identified by SDS PAGE) were concentrated separately for each protein using centricons with 10 kDa cut off.
Column 3: SEC (AKTA-prime)
Procedure (Gel Filtration - Complex formation): Samples of the gel-filtered proteins were mixed in 1:1 molar ratio and were directly applied to a S75 16/60 column equilibrated in 10 mM Hepes pH 7.5, 100 m NaCl. The flow rate was 1ml/min and the pure complex eluted at 67-77min. Concentration:The combined samples from the SEC column (identified by SDS PAGE) were concentrated using centricons with 10 kDa cut off.
Extraction
ProcedureExtraction buffer: 50 mM HEPES, pH 7.5, 500 mM NaCl, 5% Glycerol.The cell pellets (5 gr wet wt) were re-suspended in 50 ml extraction buffer containing a Protease Inhibitor Coctail tablet (Roche), and lysed in a high pressure cell disrupter. The supernatant was centrifuged for 45 minutes at 53000 g in a JA 25.5 rotor
Concentration:LigandMassSpec:LC-ESI-MStof confirmed the correct masses of 10467.0 Da (for MAP3K3B) and 11932.8 Da (for MAP2K5A) expected for these constructs after TEV treatment.
Crystallization:Crystals were grown at 4degC in 600nl sitting drops mixing 300 nanoliter of the complex (9 mg/ml in 10mM Hepes pH 7.5, 100mM NaCl ,10mM DTT taking into account the combined extinction coefficient) with 300 nl of a solution containing 0.02M CaCl 2 0.1M Acetate pH 4.6 and 30% MPD.
NMR Spectroscopy:Data Collection:Crystals were directly flash frozen in liquid nitrogen. Resolution: 2.7 Å. X-ray source: Diffraction data were collected at the Swiss Light Source (SLS) (?=0.999 Å)
Data Processing: