Rab proteins constitute the largest branch of the Ras GTPase superfamily. Rabs use the guanine nucleotide-dependent switch mechanism common to the superfamily to regulate each of the four major steps in membrane traffic: vesicle budding, vesicle delivery, vesicle tethering, and fusion of the vesicle membrane with that of the target compartment. These different tasks are carried out by a diverse collection of effector molecules that bind to specific Rabs in their GTP-bound state.
Rabs are a ubiquitously expressed family of small (20–29 kDa) monomeric Ras-like GTPases. To date, 11 Rabs have been identified in yeast (including Sec4p and the Ypt proteins) and >60 in mammalian cells. Rab proteins distinguish themselves from other members of the Ras GTPase family with five so-called Rab family sequences F1 – F5 (F1: IGVDF; F2: KLQIW (β3); F3: RFrsiT (loop 4); F4: YYRGA (α2 – loop 5); F5: LVYDIT (β4 – loop 6)). These sequences are locating in and around switch I and switch II regions, the regions that change conformation upon GDP or GTP binding.
Rab proteins have switch conformations between GTP-Rab proteins (active form) and GDP-Rab proteins (inactive form). Effector proteins interact with the active GTP-Rab proteins. Conformational differences within the switch regions (switch I and II) of different Rab proteins lead to the specific binding of various effector proteins to the targeted Rab proteins, although effectors are also found to interact with non-conserved regions of Rab proteins, which further increasing binding specificity. In some cases, the variability in Rab-GDP switch domain structures is influenced by the presence and nature of the bound metal ion. Rab proteins share some conserved residues, these residues point their side chains at different angles in the relation to the β strands of the core. These angle shifts create very distinct surfaces of related GTPases that are surely important for their ability to interact specifically with distinct effector proteins. Thus, Rab proteins have a related overall shape, but they are very different in subtle ways that will be recognized by binding partners [6].
Rab4 was found to be associated with early endosomes playing a role in their sorting and recycling. Rab4 is also implicated in the regulation of the recycling of internalized receptors back to the plasma membranes. Furthermore, Rab4 seems to have a specialized role in receptor-mediated antigen processing in B-lymphocytes, in calcium-induced a-granule secretion in platelets, and in a-amylase exocytosis in exocrine pancreatic cells. In addition, Rab4 appears to control the subcellular distribution of the glucose transporter isoform Glut4, specifically expressed in the insulin-sensitive adipose and muscle tissues. Therefore, it is assumed that Rab4 is implicated in early endosome trafficking by controlling steps necessary for sorting, recycling and degradation [reviewed in 7].
A number of proteins that specifically bind to the GTP-bound form of Rab4 have been identified, such as Rabaptin-5, Rabaptin-5b, Rabaptin-4/5a, Rabenosyn-5, Rabip4, Rabip40, Rab coupling protein_(RCP), Dynein light intermediate chain-1, syntaxin 4, and CD2AP/CMS. Most of the identified Rab effectors are divalent, interacting with more than one Rab protein. These divalent effector proteins might function as membrane-tethering molecules [reviewed in 7].
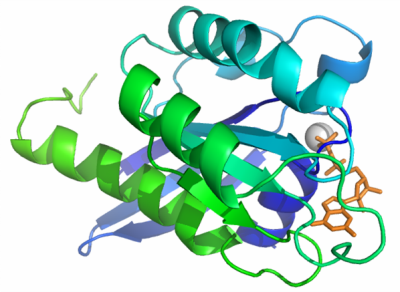
We solved the structure of Rab4B at 2.2 Å. The overall structure of this protein share common structure with other Rab proteins with relatively subtle variations. The crystal structure of Rab4B consisting of a core formed by a six stranded β-sheet surrounded by five α-helices. The conserved phosphate-binding loop (P-loop) motif in this structure is 17GXXXXGKS24, with bound Mg (II) and GDP. Superimposing this structure with crystal structure of Rab4A (2BMD), the largest deviations were observed in the switch II region (T66 – T76), the switch I loops in both protein structures are disordered. Although the sequence identity between Rab 4A and Rab 4B is high (> 80%), even in conserved secondary structure regions, exposed surfaces between two proteins are different due to the different orientations of side chains. The switch regions and the some distinct surface area may play crucial roles for the potential specific binding partner recognition.