There are multiple mechanisms used by E3's to catalyze the isopeptide bond between ubiquitin and substrates. Because ubiquitin is already activated through a thiolester bond, proximity is a common mechanism (simply bringing substrate and activated ubiquitin close to each other). The epsilon-amino group of the attacking lysine can be made more nucleophilic by catalytic residues that are in E2, E3, or the substrate (Yunus and Lima, 2006). And an oxyanion-like hole is formed by a conserved E2 asparagine, near the active site cysteine, to stabilize the tetrahedral intermediate (Passmore and Barford, 2004; Pickart and Eddins, 2004; Wu et al., 2003). Moreover, recent studies suggest that some E3's stabilize the transition state through an unknown allosteric mechanism (Ozkan et al., 2005).
The mechanism of polyubiquitin chain synthesis is poorly characterized, but likely involves a combination of the above factors. K48-based chains are commonly found in eukaryotic cells and are recognized by the proteasome. K63-based chains are less frequent and their functions are poorly understood. A subset of E2s, including the heterodimer pair MMS2/UEV1, HIP2, and CDC34, catalyze polyubiquitin chain formation. Terminal ubiquitin moieties are specifically recognized by the catalytically dead UEV1 component of the MMS2/UEV1 system. But the mechanism of K63-based and K48-based chain synthesis by HIP2 and CDC34, respectively, is unknown.
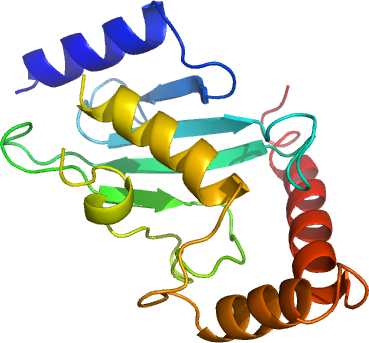
To begin elucidating the mechanism of K48-based polyubiquitin chain synthesis by human CDC34, we have solved its high resolution crystal structure. The unique 60-residue carboxy-terminal tail of CDC34, which was shown to be important for recognition by and the function of the anaphase promoting complex (Butz et al., 2005), was excluded for crystallization. CDC34 is part of a phylogenetic branch composed of its closely related human paralog, CDC34B, and the human UBC7 pair (UBC7 (2CYX) and E217K (2AWF)), all characterized by a unique stretch of acidic residues inserted immediately after the catalytic cysteine and whose function has been associated with its polyubiquitylation but not monoubiquitylation function (Petroski and Deshaies, 2005), is disordered in the current structure. Nevertheless, the structure of CDC34 has greatest structural similarity to UBC9 (1U9A) and UBC8/E220K (1YH6), which are interestingly the few E2's capable of catalyzing their reactions in an E3-independant manner. Additional experiments are needed to determine how CDC34 specifically recognizes ubiquitin’s Lys48 for the elongation reaction.