Covalent modification of histone tails plays a fundamental role in regulation of functional chromatin organization and constitutes the "histone code" critical in epigenetic regulation. Methylation of lysyl residues of histones H3 and H4 is critical to many biological processes such as heterochromatin formation, X chromosome inactivation, genome imprinting, DNA repair and transcriptional regulation. The effect of histone lysyl modification is context dependent and relies on the particular residue and the methylation state. In most instances lysyl methylation at H3K9, H3K27 and H4K20 is associated with transcriptionally silent regions, whereas methylation of H3K4, H3K36 and H3K79 appears in transcriptionally active chromatin. Methylated lysyl residues in histones recruit adaptor molecules that dictate the specific effects of methylation, e.g. methylated H3K9 associates with HP1 allowing heterochromatin formation and silencing. Furthermore, the methylation state (mono, di or tri) of Lys confers specificity to recruitment of chromatin remodeling components.
Until 2006, histone lysine methylation was considered as a stable epigenetic marker, however the discovery of enzymes capable of demethylating methylated lysyl groups dramatically changed this view. Recently two processes have been identified that catalyse demethylation of Nε-methylated lysyl residues. The FAD dependent amine oxidase LSD 1 (Lysyl demethylase 1) catalyses demethylation of mono and di-methylated, but not trimethylated lysyl residues via an imine intermediate. The recent discovery that members of the Fe(II) and 2-oxoglutarate (2OG) dependent oxygenases demethylate histone lysyl residues opened a new perspective on epigenetic regulation. To this end, catalytic activity towards methylated histone lysyl residues has been shown for members of three distinct subfamilies: JMJD1A of the JHDM2 subfamily demethylates mono and dimethylated H3K9, FBXL10 and FBXL11 of the JHDM1 subfamily catalyze demethylation of H3K36, and members of the JHDM3/JMJD2 subfamily display specificity towards tri and dimethylated states of H3K9 and H3K36.
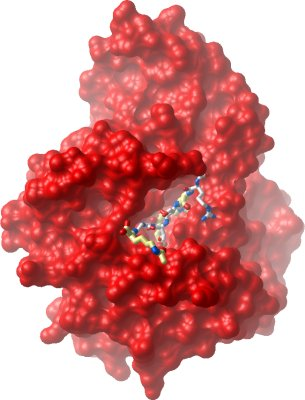
We have now determined several dead end complex structures of human JMJD2A with Ni(II) (replacing the Fe(II)), N-oxalylglycine (replacing 2OG) and different H3K9 and H3K36 substrate peptides in different methylation states.