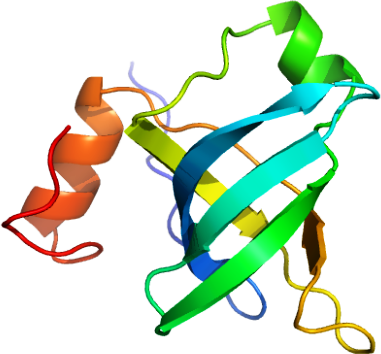
Cp-eIF1A: Cryptosporidium parvum elongation initiation fector 1A
PDB:2OQK
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:cgd8_3330
Entry Clone Source:C. parvum Iowa strain gDNA
SGC Clone Accession:CP-PF11_0447:M1-E117; plate MAC01Z:A10
Tag:N-terminal: His6-tag with integrated TEV protease site: mhhhhhhssgrenlyfq*g
Host:E. coli BL21-(DE3)-R3-pRARE2
Construct
Prelude:Sequence:gMPKNKGKGGKNRRRGKNDSEGDKRELVFKEEGQEYGQVQRMLGNGRLDAYCFDGQKRLCHIRGKMRKKVWVNPGDIVLVSLRDFQDSKGDIILKYTPDEARALKSKGEIPETTKINE
Vector:p15-TEV-LIC
Growth
Medium:M9 SeMet media kit from Medicilon
Antibiotics:100 microG/mL ampicillin and 34 microG/mL chloramphenicol
Procedure:A single colony was inoculated into 10 mL of LB with of Antibiotics and incubated with shaking at 250 rpm overnight at 37 degC. The culture was transferred into 50 mL of TB with Antibiotics in a 250 mL shaking flask and incubated at 37 degC for 3 hours. The culture was then transferred into 1.8 L of above-specified growth medium with Antibiotics and 0.3 mL of antifoam (Sigma) in a 2L bottle and cultured using the LEX system to an OD600 of ~5, cooled to 15 degC and induced with 0.5 mM isopropyl-1-thio-D-galactopyranoside (IPTG) overnight at 15 degC.
Purification
ProcedureThe cleared cell lysate was loaded onto a column containing 10 g DE-52 resin (Whatman) anion exchangeresin (previously activated with 2.5 M NaCl and equilibrated with Binding Buffer), and then directly onto a 2.5 mL Ni-NTA (Qiagen) column at approximately 1.5 mL/min. When all the lysate was loaded, both columns were washed with 15 mL binding buffer. The Ni-NTA column was then washed with 200 mL of Wash Buffer at 2 mL/min. After washing, the protein was eluted from the Ni-NTA column with 15 mL of Elution Buffer. EDTA was added immediately to 1 mM.DTT was added to 5 mM 15 minutes later.
The eluted protein was applied to a Sephadex S200 26/60 gel filtration column (GE Healthcare) pre-equilibrated with gel filtration buffer.The fractions corresponding to the eluted protein peak were collected.
The eluted protein was treated overnight at 4 degC with TEV protease, and DTT and imidazole added to 5 mM and 15 mM respectively. The protease:protein molar ratio was determined based on measured activity of the available TEV. The cleaved protein was separated from the uncleaved protein by passage through another 2.5 mL Ni-NTA column.
Five molar equivalents of GDP and MgCl2 were added to the purified protein before concentration. The buffer of the cleaved protein was exchanged against the Crystal Buffer and the protein was concentrated to 30 mg/mL using a 15 mL Amicon Ultra centrifugal filter device from Millipore (15 kD cutoff). Aliquots of the purified protein were stored at -80 degC.
Extraction
ProcedureCells were resuspended to approximately 40 mL/L of cell culture in Binding Bufferwith protease inhibitor (1 mM benzamidine-HCl and 1 mM phenylmethyl sulfonyl fluoride, PMSF). Resuspended pellets stored at -80 degC were thawed overnight at 4 degC on the day before purification. Prior to mechanical lysis, each pellet from 1 L of culture was pretreated with 0.5% CHAPS and 500 units of benzonase for 40 minutes at room temperature. Cells were mechanically lysed with a microfluidizer (Microfluidizer Processor, M-110EH) at approximately 18000 psi; and the cell lysate was centrifuged using at ~75000 x g for 20 minutes at 10 degC.
Concentration:30 mg/mL
LigandMassSpec:Crystallization:The protein was crystallized by means by sitting drop vapor diffusion in a 96-well Intelliplate. The plate was set with 0.5 microL cleaved SeMet protein (30 mg/mL) and 0.5 microL buffer in each drop, and 350 microL reservoir volume per well. Crystals grew to full size after 3 days in 2.5 M ammonium sulfate, 0.1M sodium acetate and pH 4.6 at 20 degC.
NMR Spectroscopy:Data Collection:Synchrotron: NSLS
Beam line: X8C
Wavelength: 0.97970 Å
Resolution: 1.80 Å
Data Processing:Method used: SAD
Software used: SOLVE/RESOLVE