Gastric lipases in the upper digestive tract and pancreatic lipases in the lower digestive tract hydrolyze triglycerides with short and long fatty acid chains, respectively, into monoglycerides and free fatty acids. Hydrolyzed components are absorbed by cells lining the digestive system and reassembled into large protein- and cholesterol-containing particles that then enter lymphatic ducts. There are four human versions of pancreatic lipases (PL, PLRP1, PLRP2, and the recently annotated PLRP3); although having a significant degree of sequence homology they differ in substrate specificity, behavior in bile salts, and dependence on colipase (a protein that enhances association of lipases to lipid droplets). There are also differences in temporal and spatial expression. For example, PLRP2 is expressed mainly in newborns and is less specific with respect to substrates, and PLRP2 has also been implicated as a effector of the circadian metabolic rhythm [1]. Because of their significant role in nutrient absorption, lipases are being targeted for a number of conditions related to the metabolic syndrome [2;3].
Pancreatic lipases act on the surface of emulsion particles, which are typically composed of a complex mixture of lipids. These include dietary triglycerides, proteins, oligosaccharides, polar lipids, fatty acids, cholesterol and bile salts. These micelles activate lipases by opening a two-component lid, which normally covers the active site. In the opened conformation, the lid, in conjunction with the C-terminal beta-strand subdomain, forms an extended platform for recognition of particle surfaces [4-6]. A subsite for acyl chains near the catalytic triad is created by the opened lid. Unlike PL, PLRP2 does not have a strong preference for acylglycerides over other lipids; its ability to efficiently hydrolyze galactolipids suggests that PLRP2 is dedicated to the digestion of vegetables [7-9].
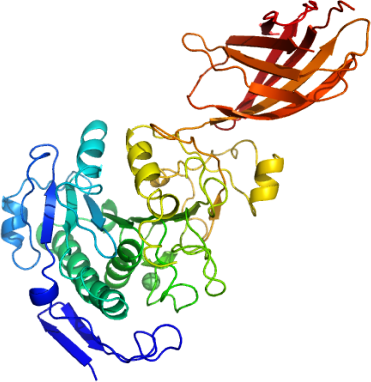
We have expressed and secreted human PLRP2 from insect cells and determined its high resolution crystal structure. The loop between beta strand 5 and alpha helix 3 (residues 94-103)is found is an open conformation; the other half of the lid (a surface loop defined by a disulfide bridge between Cys256 and Cys280) is in a conformation that closes access to substrates. Unlike other closed structures of lipases, whose lids cover and do not directly interact with catalytic residues, the lid of human PLRP2 projects a phenylalanine residue (Phe277) which fills the active site near the triad (Ser171, Asp224, and His282). This structure could help elucidate the mechanism of micellar induced activation and uncover the structural basis of substrate specificity for these medically relevant enzymes.