The HSP100 family of heat shock proteins, typically known as HSP104 in eukaryotes,
are ATP-dependent molecular chaperones with membranes including
ClpA, ClpB, ClpX and ClpY.
Specifically, ClpB is a class I heat shock protein with two nucleotide binding domains (class II HSPs have just one).
It differs from others in both function and structure [1,2].
While ClpA, ClpX and ClpY proteins are involved in protein degradation - typically working as a component of a proteolytic complex (e.g. in complex with ClpP), ClpB works with HSP70 heat shock chaperones in unfolding protein aggregates and is consequently associated with thermotolerance.
It has been speculated [2] that ClpB binds large aggregates and breaks them down into smaller aggregates.
Structurally, ClpB comprises three domains:
the N-terminal domain with double-Clp-N motif (speculated to be a protein-binding region) and two nucleotide-binding domains.
Here we present the structure of the first nucleotide binding domain (NBD1) of P. vivax ClpB protein
- specifically the region from Y168-S353.
This is an orthologue of the P. falciparum ClpB chaperone
PF08_0063
and is 86% identical in sequence.
It is also 48% identical to Thermus thermophilus ClpB (PDB ID: 1QVR),
while the Pv-ClpB-NBD1 is 67% identical to the same domain in E. coli ClpB (PDB ID: 1JBK).
Like NBD1 from E. coli ClpB (PDB ID: 1JBK),
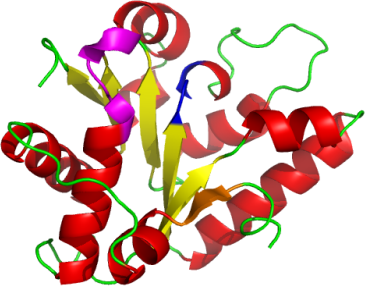
Pv-ClpB-NBD1 is an alternating α-β fold, consisting 5 parallel β-sheets, plus one extra N-terminal β-sheet (shown in orange) which runs anti-parallel to others.
This extra β strand has thus far been found only in ClpB.
Another similarity between the two structures is the lack of ATP or ADP.
Comparison against the Tt structure (1QVR) reveals that both our construct
and the Ec structure lack a C-terminal helical domain which helps to keep ATP in place.
Similar to other NBDs, our structure contains a Walker A motif
(GDPGVGKT and in magenta) which typically interacts with
a Mg2+ ion (but not found in our structure).
The DEXX Walker B motif (DEIH and in blue),
which interacts with the Walker A motif to facilitate nucleotide hydrolysis, is also present.
As found in the case of the Ec-ClpB-NBD1 [1], we were unable to co-crystallize
Pv-ClpB-NBD1 with ATP or ADP.
This is because our construct is also missing a helical domain at the C-terminus just downstream of where our construct ends.
It is clear from the T. thermophilus structure that this domain is essential to ATP binding.