Cyclosporin, also known as ciclosporin or cyclosporine, is an immunosuppressant drug originally identified from the fungus Tolypocladium inflatum Gams [1].
It is often prescribed to patients following organ transplant,
as well as for psoriassis and other auto-immune disorders.
Moreover, potent anti-parasitic activity [2] has been reported, including against malaria [3].
Concerns about adverse side-effects, however, limit the wide use of this drug for parasitic diseases.
There are over 20 variants of cyclosporin, all of which are cyclic undecapeptides.
Cyclosporin A (CsA) is best known, with other forms (e.g. B, C, D, E, G, H, etc.) varying
by individual residues or chiral deviations.
Cyclophilins are peptidyl-prolyl cis-trans isomerases (PPIase or PPI) named for their affinity for cyclosporin.
Prolyl isomerases (also PPIase) are a universally conserved family of enzymes
that converts the cis and trans isomers of peptide bonds with proline - an amino acid with a cyclic structure conferring unique conformational
rigidity to its peptide bond.
Consequently, unlkike other amino acids which preferentially form the trans peptide bond conformation,
both cis and trans isomers of proline are common.
Because proline isomerization is a rate-limiting step in protein folding, PPI's are considered chaperones.
Other PPIase groups including FK506-binding proteins (FKBP) and parvulins.
The Cryptosporidium parvum parasite is predicted to have 6 cyclophilin type PPI's encoded in its genome - cgd1_870, cgd2_4120, cgd2_1660, cgd7_520, cgd8_1560 and cgd8_2350,
in addition to two putative FK506-binding proteins - cgd6_2690 and cgd7_210.
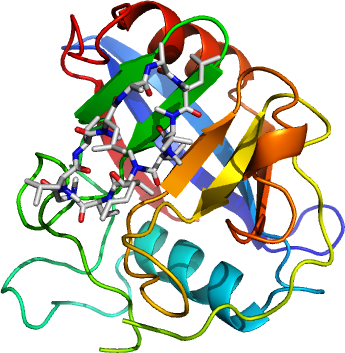
Specifically, we have obtained two crystallographic structures of the Cryptosporidium parvum cyclophilin
encoded by the gene cgd2_4120 - the apo form
(2PLU)
and the second with cyclosporin A bound
(2POY).
Like other cyclophilins, our structures do not deviate from the canonical cyclophilin fold of a barrel formed by two sets of four anti-parallel beta strands, flanked spatially (but not sequentially) by two to three helices.
The conserved CsA-interacting residues are
R78, F83, M84, Q86, G95, A124, N125, A126, Q134, F136, W145 and L146,
specifically residues 1, 2, 3, 9, 10, 11 of CsA.
The other CsA residues (4, 5, 6, 7, 8) form a composite "effector" surface with other residues on the enzyme to interact with calcineurin.
As found in the past, binding of cyclosporin does not change the structure of the enzyme itself.
Also found on the Cp cyclophilin is a
protease-like catalytic triad - H115, S122, D146.
Its functional role has yet to be determined.
See also