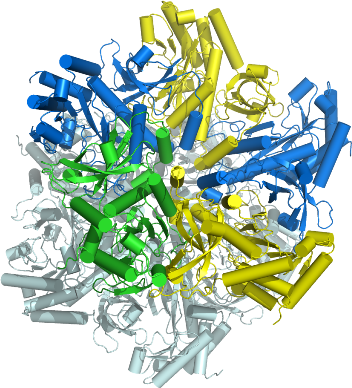
GLUL
PDB:2QC8
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:BC011852
Entry Clone Source:MGC
SGC Clone Accession:GLULA-s002
Tag:N-terminal hexahistidine tag with integrated TEV protease cleavagesite: mhhhhhhssgvdlgtenlyfq*sm
Host:BL21(DE3) gold pRARE2
Construct
Prelude:Sequence:MHHHHHHSSGVDLGTENLYFQSMLNKGIKQVYMSLPQGEKVQAMYIWIDGTGEGLRCKTRTLDSEPKCVEELPEWNFDGSSTLQSEGSNSDMYLVPAAMFRDPFRKDPNKLVLCEVFKYNRRPAETNLRHTCKRIMDMVSNQHPWFGMEQEYTLMGTDGHPFGWPSNGFPGPQGPYYCGVGADRAYGRDIVEAHYRACLYAGVKIAGTNAEVMPAQWEFQIGPCEGISMGDHLWVARFILHRVCEDFGVIATFDPKPIPGNWNGAGCHTNFSTKAMREENGLKYIEEAIEKLSKRHQYHIRAYDPKGGLDNARRLTGFHETSNINDFSAGVANRSASIRIPRTVGQEKKGYFEDRRPSANCDPFSVTEALIRTCLLNETG
Vector:pNIC-Bsa4
Growth
Medium:Antibiotics:Procedure:30 µl BL21(DE3) gold pRARE2 cells were transformed with 2 µl plasmid. The mix was kept on ice for 30 min followed by a heatshock at 42°C for 45 sec. SOC, 125 µl, was added to the cellsuspension which then was incubated for 1 hour at 37°C and plated on LA-plates containing kanamycin (50 µg/ml) and chloramphenicol (34 µg/ml). 20 ml TB (supplemented with 8 g/l glycerol, 100 µg kanamycin/ml and 34 µg/ml chloramphenicol) was inoculated with 5-10 colonies and grown overnight at 30°C. The 20 ml of the inoculation culture was added to 1.5 l TB (supplemented 8g glycerol/L and 50 µg kanamycin/ml) in 2 l bottles. The flask was incubated in the LEX system-water bath at 37 ºC until OD600 reached ~2. At this time the flask was transferred to an 18°C water bath in the LEX-system. Expression of protein was induced after approximately 1 hour by addition of 0.5 mM IPTG and continued for approximately 18 hours. Cells were harvested by centrifugation 5500 x g for 10 minutes (WCW 35.1 g).
Purification
ProcedureThe cleared lysate was loaded onto a HiTrap IMAC column (Amersham Biosciences) using an ÄktaExpress system. Eluted protein was run through a Superdex 200 16/60 gel filtration column. Fractions containing protein were pooled and the TCEP-concentration adjusted to 2 mM. Purified GS only remained stable in the presence of ATP and MnCl2 and thus 10 mM ATP and MnCl2 were added to the protein before concentration to 40 mg/ml.
Extraction
ProcedurePellets were resuspended in ~60 ml 50 mM Na-phosphate pH 7.5, 500 mM NaCl, 10 % glycerol, 10mM imidazole, 0.5 mM TCEP and Complete EDTA-free protease inhibitor (Roche Biosciences). Resuspended cells were stored at -80°C until further use. Before lyses, 8 µl of 250U/µl Benzonase (Novagen) was added to the thawed cells and sonicated (Sonics VibraCell) at 80% amplitude for 3 min (pulse: 4 s on and 4 s off).The sample was spun for 30 min at 49000 x g and the soluble fraction was decanted and filtered through a 0.45 µm syringe filter.
Concentration:LigandMassSpec:Crystallization:Crystals of the GS complex were grown using vapor diffusion at 4°C by mixing equal amounts of protein solution at 20 mg/ml including 2mM added Methionine sulfoximine and reservoir solution containing 1.1M Sodium-Malonate, 0.5% Jeffamine ED-2001, 100 mM HEPES pH 7.0. Crystals appeared after three days.
NMR Spectroscopy:Data Collection:Data to 2.6 Å resolution were collected from a single crystal at ESRF (ID29), Grenoble, France. Crystal belonged to C2 space group with cell parameters of a=181.2 Å, b=126.1 Å, c=188.2 Å, α= 90°, β=92.1°, γ= 90°.
Data Processing:The structure was solved by molecular replacement using the previously solved human Glutamine synthetase as a search model (PDB entry: 2OJW) with the program MolRep. The asymmetric unit contains a decamer. The space group was C2 with cell dimensions a=181.2Å, b=126.1Å, c=188.2Å and β=92.1°. Refmac5 was used for refinement and Coot for model building. NCS and TLS restrained refinement was used in the refinement process. The TLS groups were selected using the tlsmd server
http://skuld.bmsc.washington.edu/~tlsmd/. Data in the interval 39.9-2.60Å resolution was used and at the end of the refinement the values for R= 16.8% and Rfree= 21.7%. Coordinates for the crystal structure were deposited in the Protein Data Bank, accession code 2QC8.