Adenylosuccinate lyase (ASL) is a bi-functional enzyme which catalyzes (a) conversion of succinylaminoimidazole carboxamide ribotide to aminoimidazole carboxamide ribotide;
and (b) conversion of adenylosuccinate to adenosine 5′-monophosphate (AMP).
Fumarate is a by-product from both reactions.
As obligate scavengers of adenosine and inosine, malaria parasites are incapable of
de novo biosynthesis of purines - a trait shared by all protozoan parasites.
Accordingly, Plasmodium ASL is a mono-functional enzyme which employs the second reaction,
6-N-(1,2-dicarboxyethyl)AMP <=> fumarate + AMP
to produce AMP in the last step of the salvage pathway (shown below).
In contrast to protozoans, mammalians typically employ both de novo biosynthesis and
recycling to produce and regulate purines.
This distinction has generated interest in the Plasmodium purine salvage pathway
as a potential target of drug discovery.
By over-expressing the full-length enzyme in E. coli,
we have solved the crystallographic structure of
Plasmodium vivax adenylosuccinate lyase, or Pv-ASL
(gene ID: Pv003765).
Plasmodium vivax is the causative parasite of tertian malaria,
a typically (but not always) non-complicated and non-lethal strain of malaria occurring most frequently
in Asia.
Adenylosuccinate lyase from P. vivax protein is 60% identical to the P. falciparum orthologue
(PFB0295w,
and 40% identical ASL from E. coli.
As indicated above, the conversion of adenylosuccinate to AMP involves β-elimination of fumarate.
Enzymes which catalyze β-elimination reactions are all homo-tetramers,
including all previously solved ASL structures
(see PDB;
e.g. 1F1O,
1YIS).
The monomeric unit of Pv-ASL is shown below, along with the side view of the tetramer.
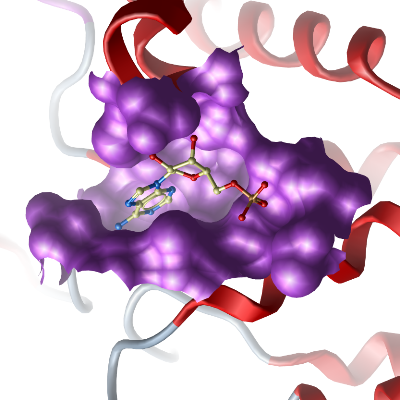
Here we present the crystal structure of Plasmodium vivax adenylosuccinate lyase (Pv003765) in complex with AMP.
Available structures for other key enzymes involved in the Plasmodium purine salvage pathway
include:
See also