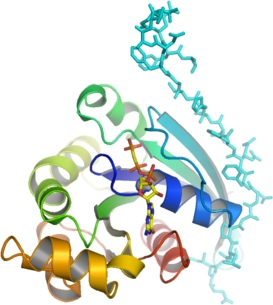
RAC3 + PAK1
PDB:2QME
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:RAC3: BC009605
Entry Clone Source:Mammalian Gene Collection
SGC Clone Accession:
Tag: N-terminal, TEV cleavable hexahistidine tag
Host:BL-21(DE3)-R3-Rosetta
Construct
Prelude:
Sequence:RAC3: smQAIKCVVVGDGAVGKTCLLISYTTNAFPGEYIPTVFDNYSANVMVDGKPVNLGLWDTAGQEDYDRLRPLSYPQTDVFLICFSLVSPASFENVRAKWYPEVRHHCPHTPILLVGTKLDLRDDKDTIERLRDKKLAPITYPQGLAMAREIGSVKYLECSALTQRGLKTVFDEAIRAVLGPAK1A CRIB domain purchased from Pepceuticals Limited : EISLPSDFEHTIHVGFDAVTGEFTGMPEQWARLLQT
Vector:pNIC28-Bsa4
Growth
Medium:
Antibiotics:
Procedure:Expression: A number of colonies from the transformation were used to innoculate 80 ml of LB media containing 50 µg/ml kanamycin and 34 µg/ml chloramphenicol, which was placed in a 37°C shaker overnight. The next day this starter culture was used to innoculate 4x 1L of TB medium (20 ml starter culture into each) containing 25 µg/ml kanamycin. When the OD600 reached ~1 the temperature was reduced to 25°C for 1 hour before induction with the addition of 1.0 mM IPTG. The expression was continued overnight (~ 16 hours).
Purification
Procedure
Column 1: IMAC. Ni-NTA (2 ml volume) in a BioRad drip column.Column 1 Buffers: Wash buffer I (WB1): 50 mM HEPES pH 7.5, 150 mM NaCl, 5 % Glycerol, 2 mM MgCl2, 10 mM Imidazole pH 8.0, 0.5 mM TCEP; Wash buffer II (WBII): 50 mM HEPES pH 7.5, 150 mM NaCl, 5 % Glycerol, 2 mM MgCl2, 30 mM Imidazole pH 8.0, 0.5 mM TCEP; Elution buffer (EB): 50 mM HEPES pH 7.5, 150 mM NaCl, 5 % Glycerol, 2 mM MgCl2, 250 mM Imidazole pH 8.0, 0.5 mM TCEP.Column 1 Procedure: 2 ml of Ni-NTA (4 ml of 50% slurry) was added to a BioRad drip column. The resin was washed with 12.5 ml of water and then 12.5 ml of WB1. The clarified cell extract was passed twice through the column. The column was then washed with 12.5 ml of WB1, and then 12.5 ml of WB2. The protein was eluted with 14 ml of EB. Nucleotide exchange procedure and TEV protease digestion: The eluted protein was concentrated to 28 mg/ml (0.5 ml volume). To this protein solution was added EDTA (to a final concentration of 20 mM - addition of 60 µl of 0.5M stock), 100 units of Calf intestinal alkaline phosphatase (CIP), DTT (to a final concentration of 1 mM), TEV protease (addition of 100 µl of homemade protease at ~ 2 mg/ml) and GppCp (addition of 200 µl of 10 mM stock). The solution was left at 4°C overnight.Column 2: Gel filtration. Hiload S200 16/60 - 120 ml volume.Column 2 Buffers: Gel Filtration buffer (GF): 50 mM HEPES pH 7.5, 100 mM NaCl, 2 mM MgCl2, 0.5 mM TCEP.Column 2 Procedure: The protein solution was loaded on the gel filtration column in GF buffer at a flow rate of 1.0 ml/min. Eluted proteins were collected in 1.75 ml fractions. The fractions containing protein were identified on a coomasie blue stained gel.Rebinding of impurities to Ni-NTA: The protein was mixed with Ni-NTA resin (pre-equilibrated into GF buffer) at 4°C for 30 minutes. The resin was spun down and the supernatent collected.
Extraction
Procedure
Procedure: The cell pellets from 4x 1L expressions were combined for purification. For each resuspended 1L pellet, 1 Complete protease inhibitor tablet (EDTA free, Roche) was dissolved in 10 ml RS and added to the cell suspension before homogenisation. The cells were lysed by 4 passes through an Emulsiflex C5 high pressure homogeniser. The cells were centrifuged for 45 mins at 16000 rpm and 4°C to remove cell debris and the cell pellet discarded.
Concentration:Concentration: The nucleotide exchanged and TEV protease cleaved RAC3A was concentrated to 13 mg/ml, distributed into 50 µl aliquots and frozen at -80°C.
Ligand
GppCp, Fragment of PAK1MassSpec:Mass spec. characterisation: Measured: 19912.1; Expected:19911.7
Crystallization:Crystallisation: Crystals grew from a 1:1 ratio of protein and precipitant solution (0.2M ammonium acetate; 0.1M citrate pH 5.6; 30% PEG 4000), using the vapour diffusion method.
NMR Spectroscopy:
Data Collection:Data Collection: Crystals were cryo-protected by equilibration into precipitant solution containing 20% ethylene glycol, and then flash frozen in liquid nitrogen. Data was collected to a resolution of 2.5 Å at the Swiss Light Source, beamline X10.
Data Processing: