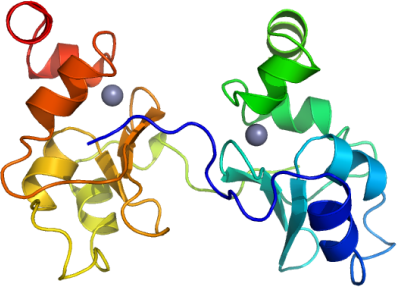
BIRC3A: Human BIR3 domain of Baculoviral Inhibitor of Apoptosis Repeat Containing 3
PDB:2UVL
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:Entry Clone Source:Mammalian Gene Collection (MGC)
SGC Clone Accession:Tag:N-terminal hexahistidine tag with integrated TEV protease cleavage site: mhhhhhhssgvdlgtenlyfq*s(m).
Host:E.coli Rosetta 2 (DE3)
Construct
Prelude:Sequence:mhhhhhhssgvdlgtenlyfqsmRYTVSNLSMQTHAARFKTFFNWPSSVLVNPEQLASAGFYYVGNSDDVKCFCCDGGLRCWESGDDPWVQHAKWFPRCEYLIRIKGQEFIRQVQAS
Vector:PNIC-BSA4
Growth
Medium:Antibiotics:Procedure:20mL TB media in 100mL shake flask, supplemented with 8 g/L glycerol, 34 µg/ml Chloramphenicol and 100 µg/mL Kanamycin, was inoculated with Rosetta 2 cells and the culture left to grow overnight at 37 °C, 175 RPM. The overnight cultures was used to inoculate fresh 1.5L TB media supplemented with 50 µg/mL kanamycin and approximately 200 µL BREOX (anti-foam solution). The large scale cultivations were grown in the LEX system at 37°C until OD600 reaches approximately 2. The flasks were then moved to 18°C and induced with 0,5 mM IPTG after 1 hour. Cultures were allowed to grow overnight at 18°C.
Purification
ProcedurePurification was performed on an ÄKTAXpress. Prior to purification, columns were equilibrated with IMAC Bind/Wash1 Buffer (HiTrap Chelating) and Gel filtration buffer (Superdex 75). The protein sample was loaded on the HiTrap Chelating column that was washed with IMAC Bind/Wash1 Buffer followed by IMAC Wash2 Buffer. Bound protein was eluted from the IMAC columns with IMAC Elution Buffer and loaded in the Gel filtration column and purified protein was slightly concentrated before TEV cleavage using Amicon Ultra 15 (Millipore) with 5000 MW cutoff. Centrifugation was performed at 15 deg in swing-out buckets, in 5 minutes intervals and maximum 15 minutes, at 3660 g.
TEV cleavage:
The HIS-tag were cleaved overnight at 4degC using HIS-taged TEV protease and the molar ratio substrate (protein):TEV-protease was 30:1. To purify the cleaved protein a matrix mix was made using Ni-agarose with GF-buffer (1:1, volumes) and 20 mM Imidazole was added to the samples and to the matrix-mix. Then 1 mL of the matrix-mix were added to the samples and incubated for 2h, at 4degC, with stirring. Finally the samples were purified, run on SDS-PAGE gel and concentrated to 16.2 g/L using vivaspin (5,000 MW CO).
Extraction
ProcedurePrepare Complete EDTA free protease inhibitor stock solution:Dissolve 12 tablets in 24 mL of IMAC binding/washing buffer.Harvest by centrifugation using the SLC-6000 rotor at 5000 rpm for 10 minutes. The cell pellet were dissolved in 2 mL Complete EDTA free protease inhibitor stock solution and resuspended in IMAC binding/washing buffer to a total volume of 90mL.
The frozen cell pellets were briefly thawed by warm water. For all samples, 4 microL (1000 U) benzonase was added to each falcon tube prior to sonication (4s/4s 3 min, 80% amplitude). The samples were diluted into a volume equal to two SA-800 tubes. (2x68 ml). The samples were centrifuged for 20 min at 20500 rpm in the Sorvall SA-800 rotor. The soluble fraction was decanted and filtered through 0.45microM filters. The samples were loaded onto the ÄKTAxpress for furher purification.
Concentration:
Ligand
MassSpec:
Crystallization:Crystals were obtained by the hanging drop vapour diffusion method using 24-well plates from Hampton Research by mixing 1uL of the protein solution (16.2 g/L) with 1microL of the well solution consisting of 16% Peg8000 and 40mM KPO4 and 20% Glycerol at room temperature.
NMR Spectroscopy:
Data Collection:X-ray data in space group P212121 (49.4, 53.7, 85.6, ? = ? = ? = 90) was collected at MaxLab (i911-3) and processed using XDS and XSCALE.
Data Processing:The structure was solved by molecular replacement with MOLREP using a BIR domain from BIRC7 (1OXN.pdb) as our search model. The initial model was then automatically rebuilt using ARP/WARP and final cycles of model building were performed in COOT with REFMAC5 as the refinement program.