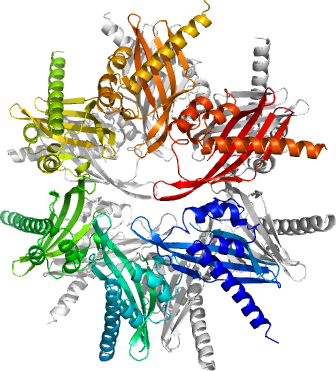
CAMK2G
PDB:2UX0
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:gi|26667203
Entry Clone Source:
SGC Clone Accession:CAMK2GA-c013
Tag:Tag sequence: *Cleavable N-terminal His 6 tag.
Host:BL21(DE3)-R3-pRARE2
Construct
Prelude:
Sequence:mhhhhhhssgvdlgtenlyfqs*SMTEDEDLKVRKQEIIKITEQLIEAINNGDFEAYTKICDPGLTSFEPEALGNLVEGMDFHKFYFENLLSKNSKPIHTTILNPHVHVIGEDAACIAYIRLTQYIDGQGRPRTSQSEETRVWHRRDGKWLNVHYHCSGAPAAPRQ
Vector:pNIC28-Bsa4.
Growth
Medium:
Antibiotics:
Procedure:Transformed 50 µl competent BL-21 (DE3) phage resistant cells with 10 µl of the plasmid DNA and plated out onto LB plate plus 50 µg/ml kanamycin. The next day colonies were picked out into fresh deep well blocks containing 1 ml TB + 50 µg/ml kanamycin which were grown overnight and glycerol stocks were prepared by adding 333 ml of 60 % glycerol to 1 ml of cell suspension, which were stored at -80°C to be used for future scale up preparations.
The glycerol stock was used to inoculate 10 ml of TB (Terrific Broth) supplemented with 50 µg/ml kanamycin. This starter culture was grown overnight at 37°C and used to inoculate a 1 liter culture in the same medium. The culture was grown at 37°C until the OD600 reached ~2.0 . After that the temperature was lowered to 18°C. Protein production was induced with 1mM IPTG and recombinant CAMK2G was expressed at that temperature over night. The next day cells were harvested by centrifugation at 4000 rpm for 15 minutes. The cell pellet was stored at -80°C degrees.
Purification
Procedure
Column 1: Ni-affinity chromatography: HisTrap FF Crude, 1 ml (GE Healthcare).
Column 2: Size exclusion chromatography HiLoad 16/60 Superdex 200
All purification steps were carried out using an AKTAexpress system (GE Healthcare) at 7ºC. The lysate was loaded on a pre-equilibrated His-trap column at 0.8 ml/min, using a standard purification method. After loading, the column was washed at 0.8 ml/min with 10 ml binding buffer, then 20 ml wash buffer, and protein was eluted with 5 ml of elution buffer. The peak fraction was collected automatically according to A280.
The CAMK2G containing fraction eluted of the Ni-affinity chromatography was automatically loaded on the SEC column at 1.2 ml/min. Eluted fractions were 95% pure as judged by SDS-PAGE.
Enzymatic treatment: The Tag sequence was cleaved by overnight treatment with recombinant, his-tagged TEV protease; the protein was passed through NiNTA beads to remove the tag and the protease.
Extraction
Procedure
The cell pellet (38 g) was re-suspended in one volume (38 ml) of 2x extraction buffer. The re-suspended cells were lysed by one passage through a Constant Systems cell breaker and subsequent sonication; the cell breaker was washed with 1x extraction buffer, bringing the total volume to 120 ml. DNA was precipitation by addition of PEI to a final concentration of 0.15 % during an incubation time of 30 min on ice, followed by a centrifugation at 17,000 rpm (4°C); The supernatant was further cleared by filtration through a 0.2 µm serum Acrodisc filter.
Concentration:12 mg/ml in SEC buffer using a centricon with a 10kDa cut off.
Ligand
MassSpec:ESI-MS revealed that the protein had the expected mass 16352.8.
Crystallization:CAMK2G was crystallized at 4 ° C using the sitting-drop vapor diffusion method . Diffraction quality crystals were obtained by mixing 75 nl of protein solution with 75 nl of 0.1M SPG pH 7.0; 60.0% MPD.
NMR Spectroscopy:
Data Collection:Crystals were flash frozen in liquid nitrogen. Diffraction data were collected to 2.7 Å at the Swiss light source beam-line X10SA at a single wavelength of 0.9999 Å.
Data Processing: