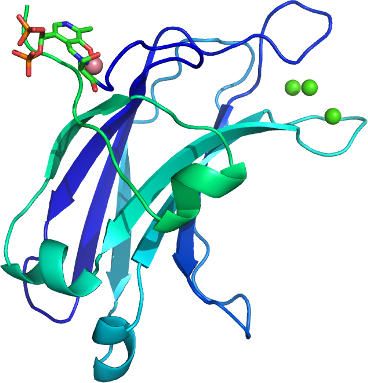
PRKCG, C2 domain
PDB:2UZP
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:gi|13384594
Entry Clone Source:Origene
SGC Clone Accession:PRKCGA-c007
Tag:Tag sequence: mhhhhhhssgvdlgtenlyfq*s(m) TEV-cleavable (*) N-terminal his6 tag.
Host:BL21(DE3)-R3
Construct
Prelude:
Sequence:mhhhhhhssgvdlgtenlyfq*smHTERRGRLQLEIRAPTADEIHVTVGEARNLIPMDPNGLSDPYVKLKLIPDPRNLTKQKTRTVKATLNPVWNETFVFNLKPGDVERRLSVEVWDWDRTSRNDFMGAMSFGVSELLKAPVDGWYKLLNQEEGEYYNVPVADADN
Vector:pNIC28-Bsa4.
Growth
Medium:LB
Antibiotics:
Procedure:Starter cultures (10 ml LB, 50 µg/ml kanamycin) were inoculated with a single colony and grown overnight. LB media (1L) was inoculated with 5 ml of culture and grown at 37°C, 170 rpm until OD600 was 0.4. The flask was then transferred to an 18°C incubator and protein expression induced with 0.1 mM IPTG (final concentration) at an OD600 of approximately 0.5. Cells were harvested the following morning by centrifugation (10min, 6500 rpm, 4°C). The pellet was resuspended in 25 ml binding buffer and frozen at -20°C.
Purification
Procedure
Column 1: DE52 anion exchange resin/IMAC Sepharose 6 Fast Flow resin, placed in series.
Column 2: Gel Filtration, HiLoad 16/60 Superdex 75 prep grade column run on an AKTA-prime system.
Procedure (Gravity feed chromatography): DE52 anion exchange resin was prepared by suspending 10 g in 100 ml of 2.5 M NaCl. The resulting gel slurry was packed in a column and allowed to settle prior to equilibrating in approximately 50 mls of Binding Buffer. Nickel charged IMAC sepharose beads were prepared by packing 4 mls of gel slurry (approximately 50% beads in 30% EtOH) in a small column and equilibrating in approximately 10 mls of Binding Buffer. Both columns were then placed in series and the protein containing supernatant passed through under gravity. This was followed by washing using 50 mls of Binding Buffer followed by 50 mls of Wash Buffer. Elution of the protein proceeded using a stepwise gradient of elution buffer: 5 mls x 30 mM imidazole, 5 mls x 50 mM imidazole and 10 mls x 250 mM imidazole.
Enzymatic treatment: Protein containing fractions were pooled and the His-tag cleaved using TEV protease (reaction proceeded overnight at 4°C).
Procedure (Gel Filtration): The protein solution was concentrated using a Vivaspin 20 ultrafiltration device with a 5 kDa MWCO to about 10 ml. A pre-equilibrated gel filtration column was injected with 5 ml of the solution and two runs undertaken at a flow rate of 0.8 ml/min.
Extraction
Procedure
The cell pellet was defrosted and cells were lysed by sonication, 10 s on, 2 s off at 40% amplitude for 8 min. The lysate was cleared by centrifugation for 45 minutes at 21,500 RPM, 4°C.
Concentration:Using Vivaspin 5 ultrafiltration spin columns with a 5 kDa MWCO the protein was dialysed against 50 mM HEPES, pH 7.5, 100 mM NaCl, 0.5 mM TCEP, 5mM Pyridoxal phosphate (PLP) and concentrated to approximately 10 mg/ml using UV absorbance at 280 nm and a predicted extinction coefficient of 27,960.
Ligand
MassSpec:Following TEV treatment of the protein mass spec confirmed the correct mass of 16,512.7 Da expected for this construct. PLP-cobalt modification during crystallization (obs, 16,798.6; exp, 16802) was confirmed by dissolving a number of crystals and subjecting to mass spec.
Crystallization:The protein was concentrated as outlined above. Prior to crystallization, 1mM Dihexanoyl- sn -Glycero-3- [Phospho-L-Serine] (sodium salt) and 10mM CaCl2 were added. Crystals were obtained at 4°C using the vapor diffusion method by mixing 180nl of the concentrated protein (10mg/ml) with 120nl of a well solution containing 1.8M ammonium sulphate, 10-15mM cobalt chloride, 0.1M MES pH6.0-7.0.
NMR Spectroscopy:
Data Collection:Crystals were cryoprotected using the well solution supplemented with an additional 25% ethylene glycol and flash frozen in liquid nitrogen. Diffraction data were collected at the SLS beam line X10SA (λ=0.999Å) to 2.0 Å.
Data Processing: