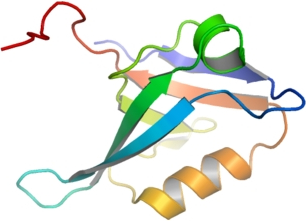
PDZD3
PDB:2V90
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:IMAGE:5186364
Entry Clone Source:MGC
SGC Clone Accession:
Tag:N-terminal, TEV cleavable hexahistidine tag
Host:BL-21(DE3)-R3-Rosetta
Construct
Prelude:
Sequence:SMKPRCLHLEKGPQGFGFLLREEKGLDGRPGQFLWEVDPGLPAKKAGMQAGDRLVAVAGESVEGLGHEETVSRIQGQGSCVSLTVVDPEADRETSV
Vector:pNIC28-Bsa4
Growth
Medium:
Antibiotics:
Procedure:Expression: 10 m l of a thawed glycerol stock was used to innoculate 40 ml of TB media containing 50 µg/ml kanamycin and 34 µg/ml chloramphenicol, which was placed in a 37°C shaker overnight. The next day this starter culture was used to innoculate 2x 1L of TB media (18 ml starter culture into each) containing 50 µg/ml kanamycin. After 5 hours, the temperature was reduced to 22°C. The incubation was continued for 1.5 hours. At OD~3, the cells were induced by the addition of 0.5 mM IPTG. The expression was continued overnight (~18 hours).
Purification
Procedure
Purification: The protein was purified using an AktaExpress system.Column 1: HisTrap 1ml.Buffers: Binding Buffer: 50 mM Hepes pH 7.4, 500mM NaCl, 5% glycerol, 5 mM Imidazole pH 7.4, 0.5 mM TCEP; Wash Buffer: 50 mM Hepes pH 7.4, 500 mM NaCl, 5% Glycerol, 25 mM Imidazole pH 7.4, 0.5 mM TCEP; Elution Buffer: 50 mM Hepes pH 7.4, 500 mM NaCl, 5% Glycerol, 250 mM Imidazole pH 7.4, 0.5 mM TCEP.Procedure: 2 ml of 50% Ni-NTA slurry was added to a clean empty 10 mm diameter gravity column (providing a resin volume of 1 ml). The resin was equilibrated with 25 ml Lysis/Binding buffer .The lysate was allowed to drip through the column twice, and the flow through was collected .The columns were washed with 12.5 ml of Lysis/Binding Buffer (Wash1) and 25 ml of Wash Buffer (Wash 2). The bound protein wash eluted with 7 x1.8 ml Elute Buffer. The 7 fractions containing protein were identified on a coomasie blue stained gel, which were pooled.Column 2: Gel filtration. Hiload S200 16/60 -120 ml volume.Buffer: Gel Filtration buffer: 50 mM Hepes pH 7.4, 500 mM NaCl, 0.5 mM TCEP.Procedure: The gel filtration column was pre-equilibrated with Gel Filtration Buffer. The Ni-NTA eluant was loaded on the gel filtration column at a flow rate of 1.2 ml/min. Eluted proteins were collected in 1.8 ml fractions. The fractions containing protein were identified on a coomasie blue stained gel.TEV protease digestion: The gel filtration fractions containing PDZD3A were pooled and 100 m l of TEV protease solution (~1 mg/ml) was added to 6 mg protein. The digestion was left overnight at 4°C.Rebinding of impurities to Ni-NTA: The protein was mixed with Ni-cellulose resin (0.3 ml, pre-equilibrated into Gel Filtration Buffer) at 4°C for 60 minutes. The resin was spun down and the supernatant was filtered through a 0.2 m M filter and collected.
Extraction
Procedure
Cell harvest: Cells were spun at 6238x g for 15 mins at 4°C. The cell pellets were placed in a -80°C freezer.Cell Lysis: Two liter-culture pellets were resuspended in lysis buffer. They were passed 4 times through an Emulsiflex C5 high-pressure homogeniser, collecting a final volume of approximately 90 ml. PEI was added to a final concentration of 0.25 % and the cell debris and precipitated DNA were spun down at 45000x g, 90 min (Beckman JA 18 17500 rpm).Lysis Buffer: 50mM HEPES pH 7.4, 500mM NaCl, 5% glycerol, 5 mM Imidazole pH 7.4, 0.5 mM TCEP, 0.2 uM PMSF
Concentration:Concentration: The TEV protease cleaved PDZD3A was concentrated to 32.4 mg/ml , distributed into 30 m l aliquots and frozen at -80°C.
Ligand
MassSpec:Mass spec. characterisation: Measured: 10244.5; Expected: 10244.4.
Crystallization:Crystallisation: Crystals grew from a 1:1 ratio of protein to precipitant solution (2M (NH4)2SO4; 0.1M BIS-TRIS pH 5.5), using the vapour diffusion method.
NMR Spectroscopy:
Data Collection:Data Collection: Resolution: 2.00 Å; X-ray source: Beamline SLS -X10SA, single wavelength.
Data Processing: