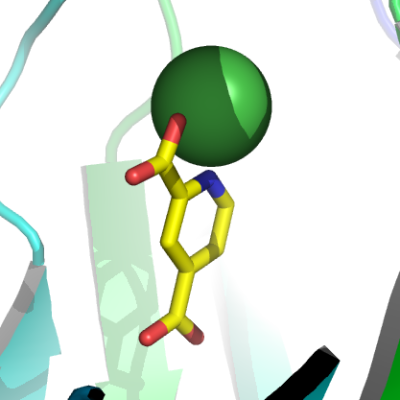
JMJD2A + inhibitor
PDB:2VD7
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:gi|7662246
Entry Clone Source:MGC
SGC Clone Accession:IMAGE:3138875
Tag:N-terminal TEV cleavable 6His tag - mhhhhhhssgvdlgtenlyfq*s(m), cleaves at *.
Host:E. coli BL21(DE3)-R3
Construct
Prelude:
Sequence:mhhhhhhssgvdlgtenlyfqsMASESETLNPSARIMTFYPTMEEFRNFSRYIAYIESQGAHRAGLAKVVPPKEWKPRASYDDIDDLVIPAPIQQLVTGQSGLFTQYNIQKKAMTVREFRKIANSDKYCTPRYSEFEELERKYWKNLTFNPPIYGADVNGTLYEKHVDEWNIGRLRTILDLVEKESGITIEGVNTPYLYFGMWKTSFAWHTEDMDLYSINYLHFGEPKSWYSVPPEHGKRLERLAKGFFPGSAQSCEAFLRHKMTLISPLMLKKYGIPFDKVTQEAGEFMITFPYGYHAGFNHGFNCAESTNFATRRWIEYGKQAVLCSCRKDMVKISMDVFVRKFQPERYKLWKAGKDNTVIDHTLPTPEAAEFLKESEL
Vector:pNIC28-Bsa4
Growth
Medium:
Antibiotics:
Procedure:Medium: TB + 50 µg/ml Kanamycin + 34 µg/ml chloramphenicol. 12 x 1 liter TB in 2.5-L baffled flasks were inoculated with 5 ml overnight culture and grown at 37°C. The protein expression was induced with 0.2 mM IPTG at OD600 = 0.8 for 18 h at 18°C. The cells were collected by centrifugation and frozen at -80°C.
Purification
Procedure
Column 1 : Ni-affinity, HisTrap FF Crude, 1 ml (GE/Amersham Biosciences)The cell extract was loaded on the column at 0.8 ml/minute on an AKTA-express system (GE/Amersham). The column was then washed with 10 volumes of lysis buffer, 50 volumes of wash buffer (lysis buffer with 40 mM imidazole), and then eluted with elution buffer (lysis buffer with 250 mM imidazole) at 0.8 ml/min. The eluted peak of A280 was automatically collected.Column 2 : Gel Filtration, Hiload 16/60 Superdex 200 prep grade, 120 ml (GE/ Amersham Biosciences)The eluted fractions from the Ni-affinity Histrap column were loaded on the gel filtration column at 1.0 ml/min. Eluted proteins were collected in 1.8 ml fractions. The target protein pool had EDTA added to a final concentration of 1 mM and was left overnight. Column 3 : HiPrep Desalting 26/10, Sephadex G-25 Fine, 57 ml (GE/ Amersham Biosciences)The protein pool from the gel filtration step was applied to the column which had been equilibrated into 50 mM tris-HCl pH 8.5, 50 mM NaCl. The resulting protein pool was 0.2 µm filtered through a cellulose acetate filter.Column 4 : 5 ml HiTrap Q Sepharose High Performance (GE/ Amersham Biosciences)The protein pool was applied to the ion exchange column and washed with column volumes of 50 mM tris-HCl pH 8.5, 50 mM NaCl. The column was eluted with a 20 column volume gradient from 50-500 mM NaCl in 50 mM tris-HCl pH 8.5, collecting 1.8 ml fractions. The fractions were analysed by SDS-PAGE and the protein pooled based on fraction purity.Column 5: HiPrep Desalting 26/10, Sephadex G-25 Fine, 57 ml (GE/ Amersham Biosciences)The protein pool from the ion exchnage step was applied to the column which had been equilibrated into 10 mM HEPES pH 7.5, 500 mM NaCl, 5% glycerol, 0.5 mM TCEP.
Extraction
Procedure
Frozen cell pellets were thawed and resuspended in a total volume of 400 ml lysis buffer. The cells were disrupted by high pressure homogenisation (15 kpsi) followed by sonication. Cell debris were removed by centrifugation for 60 minutes at 40 000 x g.
Concentration:The protein was concentrated using an Amicon Ultracel centrifugal concentrator (10 kDa MWCO) to 11 mg/ml by A280.
Ligand
2,4-pyridine dicarboxylic acidMassSpec:The mass determined for JMJD2AA-p020 was 44266 Da, in agreement with the predicted mass for the his-tagged protein.
Crystallization:Crystals were grown by vapor diffusion at 4°C. A sitting drop consisting of 100 nl protein (11 mg/ml) + 2 mM 2,4-pyridine dicarboxylic acid and 50 nl well solution was equilibrated against well solution containing 0.1 M citrate pH 5.5, 20% PEG 3350, 4 mM NiCl2. Cryo-protection of the crystals used 0.1 M citrate pH 5.5, 20% PEG 3350, 4 mM NiCl2, 20% glycerol and 5 mM 2,4-pyridine dicarboxylic acid.
NMR Spectroscopy:
Data Collection:Resolution: 2.25 Å; X-ray source: Synchrontron SLS-X10SA.
Data Processing: