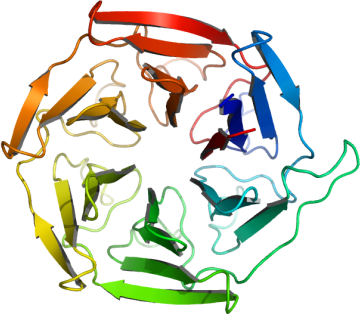
KLHL12
PDB:2VPJ
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:IMAGE:2958852
Entry Clone Source:MGC
SGC Clone Accession:
Tag:Tag sequence: mhhhhhhssgvdlgtenlyfq*s(m)TEV-cleavable (*) N-terminal his6 tag.
Host:BL21(DE3)-R3-pRARE2 (previously known as Rosetta)
Construct
Prelude:
Sequence:mhhhhhhssgvdlgtenlyfq*smQGPRTRARLGANEVLLVVGGFGSQQSPIDVVEKYDPKTQEWSFLPSITRKRRYVASVSLHDRIYVIGGYDGRSRLSSVECLDYTADEDGVWYSVAPMNVRRGLAGATTLGDMIYVSGGFDGSRRHTSMERYDPNIDQWSMLGDMQTAREGAGLVVASGVIYCLGGYDGLNILNSVEKYDPHTGHWTNVTPMATKRSGAGVALLNDHIYVVGGFDGTAHLSSVEAYNIRTDSWTTVTSMTTPRCYVGATVLRGRLYAIAGYDGNSLLSSIECYDPIIDSWEVVTSMGTQRCDAGVCVLRE
Vector: pNIC28-Bsa4
Growth
Medium:
Antibiotics:
Procedure:Starter cultures in LB media (50 ml LB, 50 µg/ml kanamycin + 34 µg/ml chloramphenicol) were inoculated from a glycerol stock and grown overnight. Four flasks containing 1L LB media were each inoculated with 10 ml of overnight culture and grown at 37°C, 160 rpm until OD600 = 0.5. The temperature was then reduced to18°C at which point protein expression was induced with 0.5 mM IPTG (final concentration). Cells were harvested the following morning by centrifugation (15 min, 5000rpm, 4°C). The pellets were each resuspended in 25 ml binding buffer and frozen at -20°C. Binding buffer: 50mM HEPES pH 7.5; 500 mM NaCl; 5 mM imidazole, 5% glycerol.
Purification
Procedure
Column 1 & 2: DE52 and IMAC Sepharose 6 Fast Flow resin, placed in series:Buffers: Binding buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 5 mM imidazole, 5% glycerol; Wash buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 30 mM Imidazole, 5% glycerol; Elution buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 50 to 250 mM Imidazole , 5% Glycerol.Column 1: Ion exchange - Nucleic acid removal. DEAE cellulose (DE52, Whatman), 10 g of resin in 2.5 x 20 cm column. The resin was hydrated in 2.5M NaCl, then washed with 20 ml binding buffer prior to loading the sample.Procedure: Supernatant was applied by gravity flow, followed by a wash with 50 ml binding buffer. The column flow-through was passed onto column 2.Column 2: Ni-affinity. Ni-sepharose (Amersham), 5 ml of 50% slurry in 1.5 x 10 cm column, washed with binding buffer.Procedure: The flowthrough from column 1 was loaded by gravity flow on the Ni-sepharose column. The column was then washed with 100 ml wash buffer under gravity flow. The protein was eluted by applying 5-ml portions of elution buffer with increasing concentration of imidazole (50 mM, 100 mM, 150 mM and 250 mM); fractions were collected until essentially all protein was eluted. 10 mM DTT was added for overnight storage together with TEV protease for cleavage of the N-terminal hexahistidine tag.Enzymatic treatment: Protein containing fractions were pooled and the His‑tag cleaved using TEV protease (reaction proceeded overnight at 4°C). Column 3: Gel Filtration, HiLoad 16/60 Superdex 200 prep grade column run on an AKTA-express system.Buffers: Gel Filtration Buffer: 50mM Hepes pH 7.5, 150mM NaCl, 0.5mM TCEP.Procedure (Gel Filtration): The protein solution was concentrated to 4ml using a 5 kD MWCO Amicon Ultra concentrator. Protein was injected onto a pre-equilibrated gel filtration column and run at a flow rate of 1.0 ml/min. Fractions containing KLHL12 were pooled.Column 4: Ni-affinity. Ni-sepharose (Amersham), 2 ml of 50% slurry in 1.0 x 10 cm column, washed with gel filtration buffer.Buffers: Elution buffers: 50 mM HEPES pH 7.5, 500 mM NaCl, 5% Glycerol with 30, 50 or 250 mM imidazole.Procedure: A further clean up step was performed to remove co-purifying proteins from IMAC. The cleaved KLHL12 from gel filtration was passed twice back through a Ni-sepharose column under gravity flow and the last flow through collected. The column was then washed with 5 ml buffer (50mM Hepes pH 7.5, 250mM NaCl), followed by 10 ml elutions with 30 mM imidazole buffer, 50 mM imidazole buffer and 250 mM imidazole buffer. All fractions were collected and run on a SDS PAGE gel. KLHL12 was found primarily in the 30 mM imidazole fraction which was pooled with the first buffer wash and the initial flow through. 10 mM DTT was added for storage.Column 5: AnionExchange Chromatography. 1 ml monoQBuffers: Buffer A: 50 mM HEPES pH 7.5; Buffer B: 50 mM HEPES pH 7.5, 1M NaClProcedure: The protein was diluted to 50 ml with BufferA and loaded onto a 1 ml monoQ anion exchange column equilibrated in the same buffer. KLHL12 was eluted with a 0 to 400 mM NaCl gradient ran over 60 ml collecting 0.75 ml fractions. KLHL12 fractions (6 ml) were combined into a 20 ml solution containing 50mM HEPES pH 7.5, 250 mM NaCl, 5mM DTT, 10mM L-Arg/10mM L-Glu.
Extraction
Procedure
Frozen pellets were thawed and fresh 1 mM PMSF and 1 mM TCEP were added. Cells were lysed by sonication. DNA was precipitated by addition of PEI to a final concentration of 0.15%, with an incubation time of 15 minutes on ice. The lysate was then centrifuged at 16,000 rpm for 60 minutes and the supernatant collected for purification.
Concentration:Protein was concentrated to 200 ml using an Amicon 10 kD MWCO Ultra concentrator then transferred to a Vivaspin 500 10 kD MWCO concentrator and concentrated further to 100 ml. The final concentration was determined by UV absorbance at 280 nm as 8.9 mg/ml using an extinction coefficient of 59820 M-1cm-1.
Ligand
MassSpec:LC- ESI -MS TOF. Expected mass (after TEV cleavage): 32873.4 Da. The expected mass was observed.
Crystallization:Crystals were obtained using the vapour diffusion method by mixing 50nl of the concentrated protein (8.9 mg/ml) with 100nl of a well solution containing the following components: 0.2M ammonium acetate, 0.1M sodium acetate pH 4.6, 30% PEG 4K. Crystallization experiments were setup at 20°C.
NMR Spectroscopy:
Data Collection:Crystals were cryo-protected using the well solution supplemented with an additional 15% PEG400 and flash frozen in liquid nitrogen. Diffraction data were collected at the SLS on beam line X10SA to 1.85 Å resolution.
Data Processing: