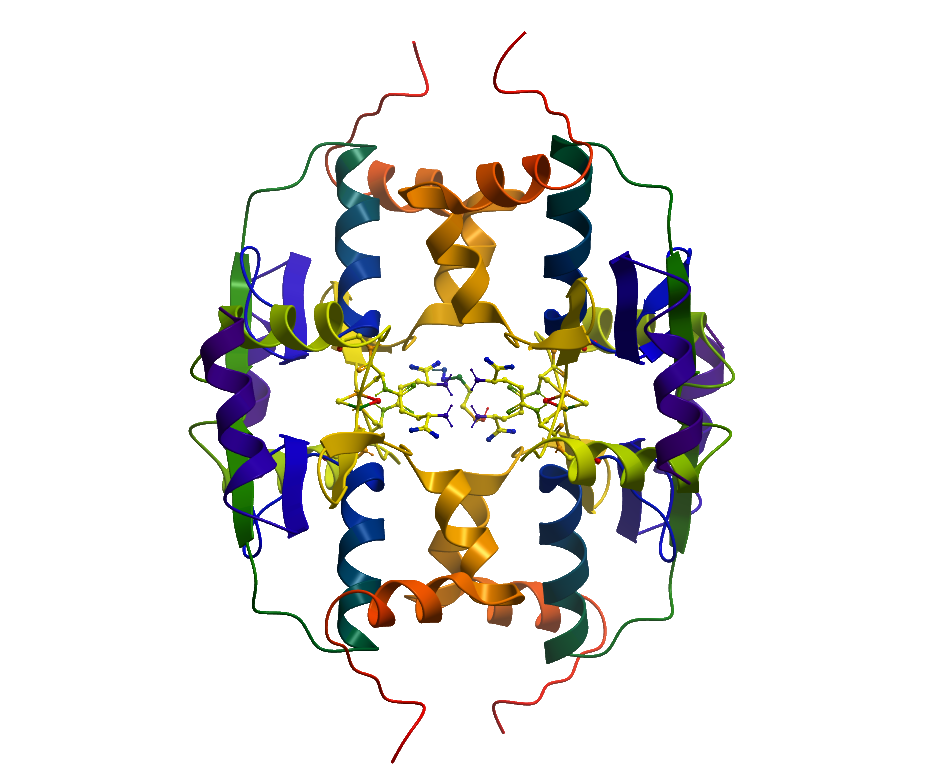
Molecular Biology
Entry Clone Accession: IMAGE:6066312
Entry Clone Source: MGC
SGC Construct ID: GLRX5A-c006
Protein Region: G35-E150
Vector: pNIC28-Bsa4
Tag: N-6HIS;N-TEV
Host: BL21(DE3)-R3-pRARE2
Sequence (with tag(s)): MHHHHHHSSGVDLGTENLYFQSMGAGGGGSAEQLDALVKKDKVVVFLKGTPEQPQCGFSNAVVQILRLHGVRDYAAYNVLDDPELRQGIKDYSNWPTIPQVYLNGEFVGGCDILLQMHQNGDLVEELKKLGIHSALLDE
Sequence after tag cleavage: SMGAGGGGSAEQLDALVKKDKVVVFLKGTPEQPQCGFSNAVVQILRLHGVRDYAAYNVLDDPELRQGIKDYSNWPTIPQVYLNGEFVGGCDILLQMHQNGDLVEELKKLGIHSALLDE
DNA Sequence: CATATGCACCATCATCATCATCATTCTTCTGGTGTAGATCTGGGTACCGAGAACCTGTACTTCCAATCCATGGGCGCGGGCGGCGGCGGCTCGGCGGAGCAGTTGGACGCGCTGGTGAAGAAGGACAAGGTGGTGGTCTTCCTCAAGGGGACGCCGGAGCAGCCCCAGTGCGGCTTCAGCAACGCCGTGGTGCAGATCCTGCGGCTGCACGGCGTCCGCGATTACGCGGCCTACAACGTGCTGGACGACCCGGAGCTCCGACAAGGCATTAAAGACTATTCCAACTGGCCCACCATCCCGCAAGTGTACCTCAATGGCGAGTTTGTAGGGGGCTGTGACATTCTTCTGCAGATGCACCAGAATGGGGACTTGGTGGAAGAACTGAAAAAGCTGGGGATCCACTCCGCCCTTTTAGATGAATGACAGTAAAGGTGGATACGGATCCGAA
Protein Expression
Medium: TB
Antibiotics: Kanamycin
Procedure: The plasmid was transformed into a phage-resistant derivative of Escherichia coli strain BL21(DE3) carrying the pRARE2 plasmid for rare codon expression. The cells were grown at 37 °C in Terrific Broth supplemented with 50 μg/ml kanamycin and 34 μg/ml chloramphenicol, until the culture reached a D600 of 1.5. The temperature was decreased to 18 °C and protein expression was induced with 0.1 mM IPTG (isopropyl β-D-thiogalactopyranoside) overnight. The cells were collected by centrifugation (5000 g for 15 min) and frozen at −80 °C.
Protein Purification
Procedure: Cell pellets were resuspended in 50 mM Hepes (pH 7.5), 500 mM NaCl, 20 mM imidazole, 5% glycerol and protease inhibitors (Complete, Sigma) and lysed by sonication (20 kHz with 2 s on and 3 s off for 10min). Cell debris and nucleic acids were removed by addition of 0.15% polyethyleneimine followed by centrifugation for 45 min at 40000 g. The proteins were purified by nickel-affinity chromatography (5 ml Ni-Sepharose FF, GE Healthcare) using a stepwise gradient of imidazole, and fractions were analysed by SDS/PAGE. The protein was concentrated and the buffer was exchanged to 50 mM Hepes (pH 7.5), 500 mM NaCl, 5% (v/v) glycerol, 10 mM GSH, 3 mM DTT (dithiothreitol) and 0.5 mM TCEP [tris-(2-carboxyethyl)phosphine] using an Amicon centrifugal filtration unit [Millipore, 3kDa MWCO (molecular mass cut-off)]. The histidine tag was removed by incubating GLRX5 with TEV protease (150 μg of TEV/10 mg of GLRX5) for 16 h at 4 °C. The protein was re-purified by applying to a nickel-affinity column (500 μl of Ni-Sepharose FF, GE Healthcare) and the flow-through was collected. The TEV-cleaved protein was concentrated using an Amicon 3K-centrifugal filtration unit and used for crystallization and further characterization.
Concentration: 93.0 mg/ml
Mass-spec Verification: Yes
Structure Determination
Crystallization: SeMet-substituted crystals were grown by vapour diffusion employing the sitting drop method, using 75 nl of protein (37 mg/ml) and 75 nl of well solution containing 70% (v/v) MPD (2-methyl-2,4-pentanediol) and 0.1 M Hepes (pH 7.5). Native crystals were grown using the same technique in a drop of 150 nl of protein (93 mg/ml) and 150 nl of well solution containing 50% (v/v) PEG [poly(ethylene glycol)] 300, 0.2 M MgCl2, 0.1 M cacodylate (pH 6.5) and 0.01 M spermine tetrahydrochloride.
Data Collection: Beamline: SLS-X10; Resolution: 2.4 Å
Data Processing: Data were processed in Mosflm, solved by anomalous scattering with SHELXD and SHELXE. The phases were improved in SHARP and Buccaneer was used to build fragments of secondary structural elements into the solvent-flattened maps. After rigid body refinement in REFMAC5 and phase improvement using Parrot, the maps enabled a first rough rebuild in Coot. The resulting model was used for molecular replacement into the higher-resolution native data with Phaser.