Entry Clone Source: MGC (S611G Mutation introduced by PCR) |
Entry Clone Accession: IMAGE:2823112 |
SGC Construct ID: RECQLA-c533 |
GenBank GI number: gi|14591904 |
Vector: pNIC-CTHF. Details [ PDF ]; Sequence [ FASTA ] or [ GenBank ]
|
Coding DNA sequence:
CTTAAGAAGGAGATATACTATGTGTTTAG
AGGATTCTGATGCCGGGGCAAGCAATGAA
TATGATTCTTCACCTGCCGCTTGGAATAA
AGAAGATTTTCCATGGTCTGGTAAAGTTA
AAGATATTCTGCAAAATGTCTTTAAACTG
GAAAAGTTCAGACCACTTCAGCTTGAAAC
TATTAACGTAACAATGGCTGGAAAGGAGG
TATTTCTTGTTATGCCTACAGGAGGTGGA
AAGAGCTTATGTTACCAGTTACCAGCATT
ATGTTCAGATGGTTTTACACTCGTCATTT
GCCCATTGATCTCTCTTATGGAAGACCAA
TTAATGGTTTTAAAACAATTAGGAATTTC
AGCAACCATGTTAAATGCTTCTAGTTCTA
AGGAGCATGTTAAATGGGTTCATGCTGAA
ATGGTAAATAAAAACTCCGAGTTAAAGCT
GATTTATGTGACTCCAGAGAAAATTGCAA
AAAGCAAAATGTTTATGTCAAGACTAGAG
AAAGCCTATGAAGCAAGGAGATTTACTCG
AATTGCTGTGGATGAAGTTCACTGCTGTA
GTCAGTGGGGACATGATTTCAGACCTGAT
TATAAGGCACTTGGTATCTTAAAGCGGCA
GTTCCCTAACGCATCACTAATTGGGCTGA
CTGCAACTGCAACAAATCACGTTTTGACG
GATGCTCAGAAAATTTTGTGCATTGAAAA
GTGTTTTACTTTTACAGCTTCTTTTAATA
GGCCAAATCTATATTATGAGGTTCGGCAG
AAGCCCTCAAACACTGAAGATTTTATTGA
GGATATTGTAAAGCTCATTAATGGGAGAT
ACAAAGGGCAATCAGGAATCATATATTGT
TTTTCTCAGAAAGACTCTGAACAAGTTAC
GGTTAGTTTGCAGAATCTGGGAATTCATG
CAGGTGCTTACCATGCCAATTTGGAGCCA
GAAGATAAGACCACAGTTCATAGAAAATG
GTCAGCCAATGAAATTCAGGTAGTAGTGG
CAACTGTTGCATTTGGTATGGGAATTGAT
AAGCCAGATGTGAGGTTTGTTATCCATCA
TTCAATGAGTAAATCCATGGAAAATTATT
ACCAAGAGAGTGGACGTGCAGGTCGAGAT
GACATGAAAGCAGACTGTATTTTGTACTA
CGGCTTTGGAGATATATTCAGAATAAGTT
CAATGGTGGTGATGGAAAATGTGGGACAG
CAGAAGCTTTATGAGATGGTATCATACTG
TCAAAACATAAGCAAATGTCGTCGTGTGT
TGATGGCTCAACATTTTGATGAAGTATGG
AACTCAGAAGCATGTAACAAAATGTGCGA
TAACTGCTGTAAAGACAGTGCATTTGAAA
GAAAGAACATAACAGAGTACTGCAGAGAT
CTAATCAAGATCCTGAAGCAGGCAGAGGA
ACTGAATGAAAAACTCACTCCATTGAAAC
TGATTGATTCTTGGATGGGAAAGGGTGCA
GCAAAACTGAGAGTAGCAGGTGTTGTGGC
TCCCACACTTCCTCGTGAAGATCTGGAGA
AGATTATTGCACACTTTCTAATACAGCAG
TATCTTAAAGAAGACTACAGTTTTACAGC
TTATGCTACCATTTCGTATTTGAAAATAG
GACCTAAAGCTAACCTTCTGAACAATGAG
GCACATGCTATTACTATGCAAGTGACAAA
GTCCACGCAGAACTCTTTCAGGGCTGAAT
CGTCTCAAACTTGTCATTCTGAACAAAGT
GATAAAAAGATGGAGGCAGAGAACCTCTA
CTTCCAATCGCACCATCATCACCACCATG
ATTACAAGGATGACGACGATAAGTGAGGA
TCC |
Tags and additions: C-terminal tag: aenlyfqshhhhhhdykddddk |
Host cells: BL21(DE3)-R3-pRARE2 |
Final protein sequence (tag sequence in lowercase):
MCLEDSDAGASNEYDSSPAAWNKEDFPWS
GKVKDILQNVFKLEKFRPLQLETINVTMA
GKEVFLVMPTGGGKSLCYQLPALCSDGFT
LVICPLISLMEDQLMVLKQLGISATMLNA
SSSKEHVKWVHAEMVNKNSELKLIYVTPE
KIAKSKMFMSRLEKAYEARRFTRIAVDEV
HCCSQWGHDFRPDYKALGILKRQFPNASL
IGLTATATNHVLTDAQKILCIEKCFTFTA
SFNRPNLYYEVRQKPSNTEDFIEDIVKLI
NGRYKGQSGIIYCFSQKDSEQVTVSLQNL
GIHAGAYHANLEPEDKTTVHRKWSANEIQ
VVVATVAFGMGIDKPDVRFVIHHSMSKSM
ENYYQESGRAGRDDMKADCILYYGFGDIF
RISSMVVMENVGQQKLYEMVSYCQNISKC
RRVLMAQHFDEVWNSEACNKMCDNCCKDS
AFERKNITEYCRDLIKILKQAEELNEKLT
PLKLIDSWMGKGAAKLRVAGVVAPTLPRE
DLEKIIAHFLIQQYLKEDYSFTAYATISY
LKIGPKANLLNNEAHAITMQVTKSTQNSF
RAESSQTCHSEQGDKKMEaenlyfqshhh
hhhdykddddk
This construct includes amino acids 49-616 of RECQL. |
Oligonucleotides used in crystallization:
5'-CGGTATTGGATCTCGACGCTCTCCCTT
5'-AGCGTCGAGATCC |
Cell growth and induction: The expression plasmid was transformed into the host strain and plated on LB-agar containing 50 µg/ml kanamycin and 35 µg/ml chloramphenicol. Several colonies were combined to inoculate a 1-ml culture in TB (+ 50 µg/ml kanamycin, 35 µg/ml chloramphenicol). The culture was grown overnight, glycerol was added to 15% v/v (from a 60% stock), and the resulting glycerol stock was frozen at -80°C in 100 µl aliquots. A loopful of cells from the glycerol stock was inoculated into 100-ml of TB medium containing 50 µg/ml kanamycin and 35 µg/ml chloramphenicol and grown overnight at 37°C.
Six cultures of 1-L each of TB medium (+ 50 µg/ml kanamycin) in 2.5L UltraYield baffled flasks were inoculated with 10 ml each of the overnight culture. The cultures were grown at 37°C until OD600 of 2-3 and then shifted to 25°C. After 30 minutes, IPTG was added to 0.1 mM, and growth continued overnight. The cells were collected by centrifugation, the pellets were scraped out and transferred to 50-ml Falcon tubes and frozen at -80°C. |
Cell extraction: Buffers: Lysis buffer: 50 mM HEPES buffer, pH 7.5, 500 mM NaCl, 10 mM imidazole, 0.5 mM TCEP, 1x Protease Inhibitors Cocktail Set VII (Calbiochem, 1/1000 dilution), and 15 units/ml Benzonase. 2x Lysis buffer contains the same components at double concentration.
Frozen cell pellets (86 g) were thawed briefly in a bath of warm water (20 - 37°C) then transferred to ice. One volume (i.e. 1 ml for every gram of cells) of 2x lysis buffer was added, followed by 1x lysis buffer to a total volume of 300-ml. The cells were resuspended by agitating and disrupted by sonication. Nucleic acids and cell debris were removed by adding 0.15% PEI (polyethyleneimine) from a 5% (w/v, pH 7.5) stock, stirring for 15 minutes, then centrifugation for 20 minutes at 25,000 xg. |
Column 1: Histrap FF 5 ml (GE Healthcare) |
Buffers:
Affinity buffer: 50 mM HEPES buffer, pH 7.5, 500 mM NaCl, 10 mM imidazole, 0.5 mM TCEP;
Wash buffer: 50 mM HEPES buffer, pH 7.5, 500 mM NaCl, 30 mM imidazole, 0.5 mM TCEP;
Elution buffer: 50 mM HEPES buffer, pH 7.5, 500 mM NaCl, 250 mM imidazole, 0.5 mM TCEP. |
Procedure: The cell extract was loaded onto the column at 5 ml/minute on an AKTA-express system (GE Healthcare). The column was then washed with 10 volumes of loading buffer, 10 volumes of wash buffer, then eluted with elution buffer at 4 ml/min. The eluted peak of A280 was automatically collected. |
Column 2: Gel filtration, HiLoad 16/60 Superdex 200 (or Superdex 75), 120 ml. |
Buffer: GF buffer: 10 mM HEPES, pH 7.5, 500 mM NaCl, 5% glycerol, 0.5 mM TCEP. |
Procedure: The eluted fraction was loaded and fractionated on the gel filtration column in GF buffer at 1.2 ml/min. 2-ml fractions were collected at the A280 peaks. The fractions were analyzed by SDS-PAGE and relevant fractions were pooled. The protein was concentrated to 10 mg/ml using a centricon centrifugal device (30 kDa MWCO). |
Protein characterization: Mass spectrometry characterization: Observed mass 67222, calculated mass 67294; (unexplained mass difference of -72). This may partly arise from cleavage of the N-terminal methionine.
The protein migrated in gel filtration with a size corresponding to a dimer (130 kDa). |
Crystallization: The two DNA oligonucleotides were mixed at a final concentration of 100 µM each in 10 mM HEPES, pH 7.5, 50 mM NaCl and annealed by heating to 95°C and gradual cooling to 15°C. To form protein:DNA complexes, the protein and DNA were combined and diluted to concentrations of 3.3 µM and 4 µM, respectively, in a buffer consisting of 10 mM HEPES, pH 7.5 and 10 mM NaCl. The complexes were then concentrated by ultrafiltration to approximately 140 µM protein.
Crystals were grown by vapour diffusion from nanolitre sitting drops at 4°C. The best crystals were obtained from drops comprising 150nl protein:DNA solution (10mg/ml) and 50nl reservoir solution (0.2M sodium sulphate, 20% PEG3350, 10% ethylene glycol) equilibrated against reservoir solution. |
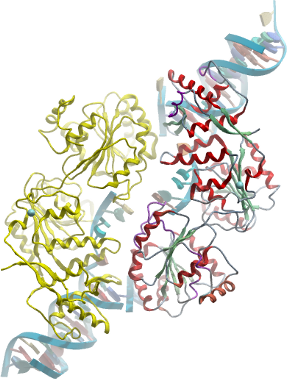
Data collection / Structure determination: Diffraction data were collected to a resolution of 2.9Å on beamline I04 at the Diamond Light Source (Harwell, UK) from a crystal vitrified in reservoir solution supplemented with 25% ethylene glycol. Crystals belong to spacegroup P21 and contain a single RecQ dimer associated with two DNA duplexes. The structure was solved by molecular replacement using the coordinates of the unbound enzyme (2V1X) as a search model using the program PHASER. The complex has been refined with PHENIX using appropriate restraints (TLS, grouped B, NCS). |