Entry Clone Source: Site directed mutagenesis |
Entry Clone Accession: n/a |
SGC Construct ID: LHPPA-c503 |
GenBank GI number: gi|33636766 |
Vector: pNIC28-Bsa4. Details [PDF ]; Sequence [ FASTA ] or [ GenBank ]
|
Coding DNA sequence:
CATATGCACCATCATCATCATCATTCTTCT
GGTGTAGATCTGGGTACCGAGAACCTGTAC
TTCCAATCCATGGCACCGTGGGGCAAGCGG
CTGGCTGGCGTGCGCGGGGTGCTGCTTGAC
ATCTCGGGCGTGCTGTACGACAGCGGCGCG
GGCGGCGGCACGGCCATCGCCGGCTCGGTG
GAGGCGGTGGCCAGACTGAAGCGTTCCCGG
CTGAAGGTGAGGTTCTGCACCAACGAGTCG
GCCGCCTCCCGGGCAGAGCTGGTGGGGCAG
CTTCAGAGGCTGGGATTTGACATCTCTGAG
CAGGAGGTGACCGCCCCGGCACCAGCTGCC
TGCCAGATCCTGAAGGAGCGAGGCCTGCGA
CCATACCTGCTCATCCATGACGGAGTCCGC
TCAGAATTTGATCAGATCGACACATCCAAC
CCAAACTGTGTGGTAATTGCAGACGCAGGA
GAAAGCTTTTCTTATCAAAACATGAATAAC
GCCTTCCAGGTGCTCATGGAGCTGGAAAAA
CCTGTGCTCATATCACTGGGAAAAGGGCGT
TACTACGCTGCCACCTCTGGCCTGATGCTG
GACGTTGGTCCCTACATGAAGGCGCTTGAG
TATGCCTGTGGCATCAAAGCCGAGGTGGTG
GGGAAGCCTTCTCCTGAGTTTTTCAAGTCT
GCCCTGCAAGCGATAGGAGTGGAAGCCCAC
CAGGCCGTCATGATTGGGGACGATATCGTG
GGCGACGTCGGCGGTGCCCAGCGGTGTGGA
ATGAGAGCGCTGCAGGTGCGCACCGGGAAG
TTCAGGCCCAGTGACGAGCACCATCCGGAA
GTGAAGGCTGATGGGTACGTGGACAACCTC
GCAGAGGCAGTGGACCTGCTGCTGCAGCAC
GCCGACAAGTGACAGTAAAGGTGGATACGG
ATCCGAA
|
Tags and additions: N-terminal, TEV cleavable hexahistidine tag |
Final protein sequence:
SMAPWGKRLAGVRGVLLDISGVLYDSGAGG
GTAIAGSVEAVARLKRSRLKVRFCTNESAA
SRAELVGQLQRLGFDISEQEVTAPAPAACQ
ILKERGLRPYLLIHDGVRSEFDQIDTSNPN
CVVIADAGESFSYQNMNNAFQVLMELEKPV
LISLGKGRYYAATSGLMLDVGPYMKALEYA
CGIKAEVVGKPSPEFFKSALQAIGVEAHQA
VMIGDDIVGDVGGAQRCGMRALQVRTGKFR
PSDEHHPEVKADGYVDNLAEAVDLLLQHAD
K
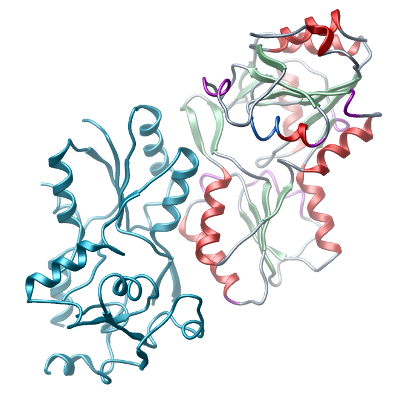
This sequence represents an engineered construct where high-entropy side-chains from two loop regions (Q58, K59, K160, E161) were mutated to alanine (residues underlined) to improve crystallization. |
Expression strain: BL-21(DE3)-R3-Rosetta (A homemade phage resistant version of BL21(DE3) containing the pRARE2 plasmid from Rosetta II (DE3) cells). |
Transformation: The construct DNA was transformed into competent cells of the expression strain by a standard heat shock procedure. |
Glycerol stock preparation: A number of colonies were used to inoculate 1 ml of LB media containing 50 µg/ml kanamycin and 34 µg/ml chloramphenicol, which was incubated in a 37°C shaker overnight. The next day glycerol stocks were prepared from this overnight culture. |
Expression: 10 µl of a glycerol stock was used to inoculate 40 ml of LB containing 50 µg/ml kanamycin and 34 µg/ml chloramphenicol, which was incubated at 37°C overnight. The next day, 8ml of the starter culture was used per litre TB containing 50 µg/ml kanamycin. The culture was incubated at 37°C until OD~1, at which point the it was induced with 0.1mM IPTG and expression was continued overnight at 18°C/170 rpm. |
Cell harvest: Cells were pelleted at 6238x g for 15 min at 4°C, and stored at -80°C. The yield was 7g cells / litre culture. |
Cell Lysis:
Lysis Buffer: 50mM HEPES pH 7.4, 500mM NaCl, 5% glycerol, 10 mM Imidazole pH 7.4, 0.5 mM TCEP, 1mM PMSF
The pellets were resuspended in lysis buffer. They were passed 4 times through an Emulsiflex C5 high-pressure homogeniser, collecting a final volume of approximately 20 ml/ litre culture. 0.25 % PEI (v/v) was added and cell debris and precipitated DNA were spun down at 45000x g, 60 min ( Beckman JA 18 17500 rpm). The supernatant was collected to which 12.5 U of Benzonase was added with a 30-60 min incubation on ice. |
Purification:
Wash Buffer: 50 mM Hepes pH 7.4, 500 mM NaCl, 5% Glycerol, 30 mM Imidazole pH 7.4, 0.5 mM TCEP
Elution Buffer: 50 mM Hepes pH 7.4, 500 mM NaCl, 5% Glycerol, 250 mM Imidazole pH 7.4, 0.5 mM TCEP
Gel Filtration buffer: 50 mM Hepes pH 7.4, 500 mM NaCl, 5% glycerol, 0.5 mM TCEP
Purification was performed in an Akta Express system (GE Healthcare) with an automated program for Ni-affinity (HisTrap FF) and gel filtration (HiLoad 16/60 S200) chromatography steps.
Step 1 Ni-affinity, HisTrap Crude FF, 1 ml: The clarified cell extract was loaded on the column at 0.8 ml/min on the AKTA-express system. The column was washed with 20 cv of binding buffer, 10 cv of wash buffer, and then eluted with elution buffer at 0.8 ml/min. The eluted peak of A280 was automatically collected.
Step 2 Gel filtration, Hiload 16/60 Superdex S200 prep grade, 120 ml: The eluted fractions from the Ni-affinity column was loaded on the gel filtration column pre-equilibrated in GF buffer at 1.2 ml/min. Eluted proteins were collected in 1.8 ml fractions and analyzed on SDS-PAGE. |
TEV protease digestion: Peak fractions containing LHPPA were pooled and TEV protease was added at a molar ratio of 1:30. The digestion was left overnight at 4 °C. Mass spec confirmed complete digestion. His-TEV and contaminating proteins were removed by passing to 300µl Ni resin pre-equilibrated in GF buffer. The flow through containing TEV-cleaved protein was collected, concentrated to 27 mg/ml using a centricon centrifugal device with a 10kDa MWCO, frozen at -80°C in 70µl aliquots. |
Mass spectrometry characterization:
Measured: 29051.8
Expected: 29051.3
|
Crystallization:
Crystals were grown at 4°C by vapour diffusion in sitting drops mixing protein (27mg/ml) and precipitant solution (0.3M Na formate and 25% (w/v) PEG3350) in a 2:1 ratio. Crystals were cryo-protected in 25% glycerol and flash frozen in liquid nitrogen.
|
Data Collection: Data was collected to a resolution of 1.92 Å at beamline I02 of Diamond Light Source. |