Entry Clone Source: MGC |
Entry Clone Accession: IMAGE:3686411 |
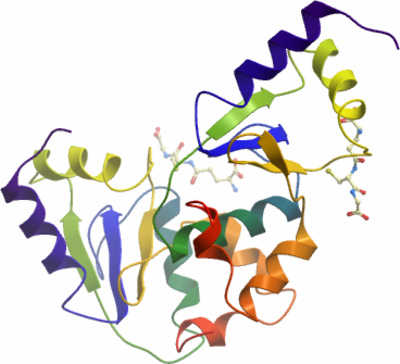
SGC Construct ID: TXNL2A-c020 |
GenBank GI number: gi|95113651 |
Vector: pNIC28-Bsa4. Details [PDF ]; Sequence [ FASTA ] or [ GenBank ] |
Amplified construct sequence:
CATATGCACCATCATCATCATCATTC
TTCTGGTGTAGATCTGGGTACCGAGA
ACCTGTACTTCCAATCCATGGCTCCC
AAATTAGAGGAAAGGCTCAAAGTGCT
GACAAATAAAGCTTCTGTGATGCTCT
TTATGAAAGGAAACAAACAGGAAGCA
AAATGTGGATTCAGCAAACAAATTCT
GGAAATACTAAATAGTACTGGTGTTG
AATATGAAACATTCGATATATTGGAG
GATGAAGAAGTTCGGCAAGGATTAAA
AGCTTACTCAAATTGGCCAACATACC
CTCAGCTGTATGTGAAAGGGGAGCTG
GTGGGAGGATTGGATATTGTGAAGGA
ACTGAAAGAAAATGGTGAATTGCTGC
CTATACTGAGAGGAGAAAATTGACAG
TAAAGGTGGATACGGATCCGAA
|
Final protein sequence (Tag sequence in lowercase):
mhhhhhhssgvdlgtenlyfq^smAP
KLEERLKVLTNKASVMLFMKGNKQEA
KCGFSKQILEILNSTGVEYETFDILE
DEEVRQGLKAYSNWPTYPQLYVKGEL
VGGLDIVKELKENGELLPILRGEN
^ TEV cleavage site |
Tags and additions: Cleavable N-terminal His6 tag. |
Host: BL21 (DE3)R3-pRARE2 (Phage resistant strain).
|
Growth medium, induction protocol: TB supplemented with 50µg/ml kanamycin and 34µg/ml chloramphenicol. 4L TB in two 4L flask were inoculated with 40ml overnight culture and grown at 37°C unti OD600 reached 2.5. The temperature was then decreased to 18°C and the protein expression induced with 0.1 mM IPTG overnight. The cells were collected by centrifugation at 4000rpm for 30 minutes and frozen at -80°C.
Lysis buffer: 50 mM HEPES, pH 7.5; 500 mM NaCl; 10 mM Imidazole; 5% Glycerol; 0.5 mM TCEP; 1 mM PMSF; 3 U/ml Benzonase.
Extraction buffer, extraction method: The cell pellet was resuspended in a total volume of 200ml lysis buffer and the cells disrupted by sonication. Nucleic acid and cell debris were removed by adding 0.15% PEI, followed by centrifugation for 60 minutes at 40,000g. The supernatant was further clarified by filtration (0.20µm). |
Column 1: Ni-sepharose, HisTrap FF, 5ml (GE healthcare). |
Column 1 Buffers:
Binding buffer: 50 mM HEPES, pH 8.0; 200 mM NaCl; 10 mM Imidazole; 0.5 mM TCEP; 5 mM GSH; 5 U/ml of Benzonase and 1 tablet complete protease inhibitors per 50ml.
Wash buffer: 10 mM HEPES, pH 7.5; 200 mM NaCl; 20 mM Imidazole; 0.5 mM TCEP; 5 mM GSH.
Elution buffer: 10 mM HEPES, pH 7.5; 200 mM NaCl; 250 mM Imidazole; 0.5 mM TCEP; 5 mM GSH.
Buffer exchange: 10 mM HEPES, pH 7.5; 200 mM NaCl; 10 mM GSH; 1 mM TCEP. |
Column 1 Procedure: The cell extracted was applied onto a 5ml Ni-sepharose FF gravity column equilibrated with lysis buffer. The column was subsequently washed with 10 volumes of lysis buffer followed by 10 wash buffer. The protein was eluted by 2x3 volumes of elution buffer and the buffer exchanged to 10 mM HEPES pH 7.5, 200 mM NaCl, 10 mM GSH, 1 mM TCEP. All fractions were collected and analysed by SDS-PAGE. The protein had visible dark brown colour and absorbed light at 410nm and 510nm when analysed by UV-visible spectroscopy. |
Protein concentration: The protein was concentrated in Amicon 3K to 102mg/ml. The protein concentration was determined spectrophotometically using the predicted molar extinction coefficient 12950 (M-1cm-1) and the predicted mass of 14472.5Da. |
Mass spectrometry characterization: The mass determined for TXNL2Ap031 was 14472.5Da, in agreement with the predicted mass of the protein with an N-terminal histidine tag. |
Crystallisation: TXNL2A was crystallised by vapour diffusion at 4°C from a sitting drop consisting of 100nl protein (25mg/ml) and 50nl 0.1 Citrate pH 3.5, 25% PEG3350. The crystal was transferred to a cryo protectant composed of 25% ethylene glycol before flash-cooling in liquid nitrogen. |
Data collection:
Resolution: 2.1Å.
X-ray source: Diamond Light Source beamline I03. |