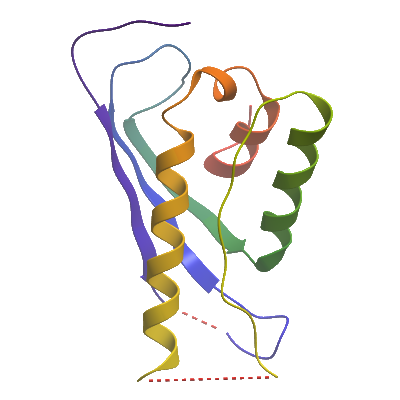
SNX3A (2YPS) Materials & Methods |
Entry clone source: MGC |
Entry Clone Accession: BC014580 |
SGC Construct ID: SNX3A-c018 |
Vector: pNIC28-Bsa4. Details [PDF ]; Sequence [ FASTA ] or [ GenBank ]. |
DNA sequence:
ATGGCGGAGACCGTGGCTGACACCCG
GCGGCTGATCACCAAGCCGCAGAACC
TGAATGACGCCTACGGACCCCCCAGC
AACTTCCTCGAGATCGATGTGAGCAA
CCCGCAAACGGTGGGGGTCGGCCGGG
GCCGCTTCACCACTTACGAAATCAGG
GTCAAGACAAATCTTCCTATTTTCAA
GCTGAAAGAATCTACTGTTAGAAGAA
GATACAGTGACTTTGAATGGCTGCGA
AGTGAATTAGAAAGAGAGAGCAAGGT
CGTAGTTCCCCCGCTCCCTGGGAAAG
CGTTTTTGCGTCAGCTTCCTTTTAGA
GGAGATGATGGAATATTTGATGACAA
TTTTATTGAGGAAAGAAAACAAGGGC
TGGAGCAGTTTATAAACAAGGTCGCT
GGTCATCCTCTGGCACAGAACGAACG
TTGTCTTCACATGTTTTTACAAGATG
AAATAATAGATAAAAGCTATACTCCA
TCTAAAATAAGACATGCCTGA
|
Final protein sequence (His6 affinity Tag sequence in lowercase):
mhhhhhhssgvdlgtenlyfq^smPP
SNFLEIDVSNPQTVGVGRGRFTTYEI
RVKTNLPIFKLKESTVRRRYSDFEWL
RSELERESKVVVPPLPGKAFLRQLPF
RGDDGIFDDNFIEERKQGLEQFINKV
AGHPLAQNERCLHMFLQDEIIDKSYT
^ TEV protease recognition site
|
Tags and additions: Cleavable N-terminal His6 tag |
Host: BL21(DE3)-R3-pRARE2. Phage-resistant strain. |
Expression: A glycerol stock was used to inoculate 2X60 ml of TB media containing 50 ìg/ml kanamycin and 50 ìg/ml chloramphenicol, which was placed in a 37°C shaker overnight. The next day this starter culture was used to inoculate 12L of TB media (10 ml starter culture used per 1L) containing 50 ìg/ml kanamycin. When the OD600 reached approximately 1.0 the temperature was reduced to 18°C and after a further 30 minutes the cells were induced by the addition of 0.1 mM IPTG. Expression was continued overnight.
|
Cell harvest: Cells were harvested by centrifugation at 16,000 RPM after which the supernatant was poured out and the cell pellet either placed in a -80°C freezer or used directly for purification.
|
Cell Lysis: Cell pellets were dissolved in approximately 50ml lysis buffer and broken by passing through the homogeniser (x6) at a constant pressure of 15KPa. The cell debris was pelleted at 16,000 RPM and the supernatant used for further purification.
Binding buffer: 50 mM Hepes (pH 7.5), 500 mM NaCl, 5% Glycerol, 20 mM Imidazole pH 7.5, 0.01mM TCEP
|
Column 1: Ni-NTA (5.0 ml volume in a gravity-flow column). |
Column 1 Buffers:
Binding buffer: 50 mM Hepes (pH 7.5), 500 mM NaCl, 5% Glycerol, 20 mM Imidazole pH 7.5, 0.01mM TCEP
Wash buffer: 50 mM Hepes (pH 7.5), 500 mM NaCl, 5% Glycerol, 40 mM Imidazole pH 7.5, 0.01mM TCEP
Elution buffer: 50 mM Hepes (pH 7.5), 500 mM NaCl, 5% Glycerol, 250 mM Imidazole pH 7.5, 0.01mM TCEP
|
Column 1 Procedure: The clarified cell extract was incubated with 5.0 ml pre-equilibriated 50% Ni_NTA bead solution for 1 hour at 4°C with rotation after which it was passed through a glass column. The column was then washed with 50ml Binding Buffer (2 x 25ml) and 50 ml Wash Buffer (2 x25 ml). The protein was eluted with 50 ml of Elution Buffer in 5 x 5 ml fractions.
|
Column 2: Superdex s200 16/60 Gel Filtration. |
Column 2 Buffers:
Gel Filtration buffer: 10 mM Hepes (pH 7.5), 500 mM NaCl, 5% Glycerol, 0.01mM TCEP
|
Column 2 Procedure: Elution fraction 1 and 2 were then pooled and concentrated to 5 ml (10 kDa mwco concentrator) and applied to the GF column (pre-equlibriated in GF buffer) at 1.0 ml/min. 1.0 ml fractions were collected.
|
Enzymatic treatment: The N-terminal His6- tag was cleaved by incubating overnight with TEV (20°C). Cleaved protein was purified by batch binding on 1ml pre-equilibriated 50% Ni-NTA bead solution. The column was then washed with 2x1ml Gel Filtration buffer, 2x1ml Wash buffer. |
Concentration: To set up plates the sample was concentrated to 15.87 mg/ml using a 10 kDa mwco concentrator.
|
Mass spec characterization:
Expected mass: 15360.8 Da, Measured mass: 15630.781 Da
|
Crystallization: Crystals were grown by vapour diffusion in sitting drop at 20°C. A sitting drop consisting of 75 nl protein and 75nl well solution was equilibrated against well solution containing 3M formate. |
Data Collection: Resolution: 2.60 Å
X-ray source: Swiss Light Source beamline IO3
|