ZMPSTE24A (2YPT) Materials & Methods |
Entry Clone Source: MGC |
Entry Clone Accession: IMAGE: 5269064 |
SGC Construct ID: ZMPSTE24A-c029 |
GI Number: 10269 |
Vector: pFB-CT10HF-LIC. Details [PDF ]; Sequence [ FASTA ] or [ GenBank ] |
DNA sequence:
CTTAAGAAGGAGATATACTATGGGGA
TGTGGGCATCGCTGGACGCTTTGTGG
GAGATGCCGGCCGAGAAGCGTATCTT
CGGGGCCGTGCTGCTCTTTTCCTGGA
CAGTGTATCTTTGGGAGACCTTCCTA
GCACAGCGGCAGAGAAGGATATATAA
AACAACAACTCATGTACCACCGGAGT
TAGGACAGATCATGGATTCTGAAACA
TTTGAGAAATCTCGACTCTATCAACT
GGATAAAAGCACTTTCAGCTTCTGGT
CAGGACTCTATTCAGAGACTGAAGGC
ACTCTTATTCTTCTCTTTGGAGGAAT
ACCTTATCTCTGGAGACTTTCTGGAC
GGTTCTGTGGTTATGCTGGCTTTGGA
CCAGAATATGAGATCACTCAGTCCCT
GGTGTTTCTGCTGTTGGCTACACTTT
TCAGTGCATTGGCTGGTTTGCCATGG
AGTCTTTATAATACTTTTGTGATAGA
AGAAAAACATGGCTTCAATCAACAGA
CTTTGGGGTTCTTCATGAAAGATGCA
ATCAAGAAATTTGTTGTGACTCAGTG
CATTTTGTTGCCTGTGTCTTCACTTC
TACTTTACATTATTAAAATTGGGGGT
GACTATTTTTTTATTTATGCCTGGCT
GTTCACATTAGTTGTGTCTCTGGTTC
TTGTCACAATCTATGCTGATTATATT
GCCCCTTTATTTGACAAATTCACACC
TCTGCCTGAGGGAAAGCTTAAAGAAG
AAATTGAAGTAATGGCAAAGAGTATT
GACTTTCCTTTGACGAAGGTGTATGT
TGTGGAAGGATCTAAACGCTCTTCCC
ACAGCAATGCTTATTTTTATGGCTTC
TTCAAGAACAAGCGAATAGTTTTGTT
TGACACTCTACTAGAAGAGTACTCTG
TACTAAACAAAGACATCCAGGAGGAT
TCTGGCATGGAACCCCGCAATGAGGA
AGAAGGGAACAGTGAAGAAATAAAAG
CTAAAGTTAAAAATAAGAAACAAGGA
TGTAAAAATGAGGAGGTACTCGCTGT
ACTAGGCCATGCACTGGGGCACTGGA
AGTTGGGACATACAGTCAAAAATATC
ATTATTAGCCAGATGAATTCTTTCCT
GTGTTTTTTTTTATTTGCTGTATTAA
TTGGTCGAAAGGAGCTTTTTGCTGCA
TTTGGTTTTTATGATAGCCAACCCAC
TCTTATTGGACTATTGATCATCTTCC
AGTTTATTTTTTCACCTTACAATGAG
GTTCTTTCTTTTTGCCTAACAGTCCT
AAGCCGCAGATTTGAGTTTCAAGCTG
ATGCATTTGCCAAGAAACTTGGGAAG
GCTAAAGACTTATATTCTGCTTTAAT
CAAACTTAACAAAGATAACTTGGGAT
TCCCTGTTTCTGACTGGTTGTTCTCA
ATGTGGCATTATTCTCATCCTCCACT
GCTAGAGAGACTTCAAGCTTTGAAAA
CTATGAAGCAACACGCAGAGAACCTC
TACTTCCAATCGCACCATCATCACCA
TCACCATCACCACCATGATTACAAGG
ATGACGACGATAAGTGAGGATCC
|
Expressed sequence (small letters refer to tag sequence):
MGMWASLDALWEMPAEKRIFGAVLLF
SWTVYLWETFLAQRQRRIYKTTTHVP
PELGQIMDSETFEKSRLYQLDKSTFS
FWSGLYSETEGTLILLFGGIPYLWRL
SGRFCGYAGFGPEYEITQSLVFLLLA
TLFSALAGLPWSLYNTFVIEEKHGFN
QQTLGFFMKDAIKKFVVTQCILLPVS
SLLLYIIKIGGDYFFIYAWLFTLVVS
LVLVTIYADYIAPLFDKFTPLPEGKL
KEEIEVMAKSIDFPLTKVYVVEGSKR
SSHSNAYFYGFFKNKRIVLFDTLLEE
YSVLNKDIQEDSGMEPRNEEEGNSEE
IKAKVKNKKQGCKNEEVLAVLGHALG
HWKLGHTVKNIIISQMNSFLCFFLFA
VLIGRKELFAAFGFYDSQPTLIGLLI
IFQFIFSPYNEVLSFCLTVLSRRFEF
QADAFAKKLGKAKDLYSALIKLNKDN
LGFPVSDWLFSMWHYSHPPLLERLQA
LKTMKQHAENLYFQ^shhhhhhhhhh
dykddddk
|
Tags and additions: C-terminal, TEV cleavable decahistidine / FLAG tag. |
Host: Spodoptera frugiperda (SF9) insect cells |
Growth Medium & Induction Protocol: Insect cells with a density of 2x106 per litre of cell culture in SF900 medium (Invitrogen) were infected with recombinant baculovirus (5ml P2 virus per litre cell culture) and incubated for 72 hours. Cells were harvested by centrifugation and the pellet was flash frozen in liquid N2.
|
Extraction buffer, extraction method: Frozen pellets were thawed and re-suspended in extraction buffer supplemented with 1% OGNG/ 0.1% CHS and incubated at 4°C on a tube rotator. The extracted protein was separated from insoluble membranes by centrifugation and collected for purification.
|
Extraction/wash buffer: 50 mM HEPES, pH 7.5; 200 mM NaCl |
Column 1: Co-affinity. Cobalt Talon (Clontech), 0.75 ml of 50 % slurry/ 1L of cells in 1.5 x 10 cm column, washed with extraction/ wash buffer. |
Column 1 Buffers:
Extraction/wash buffer: 50 mM HEPES, pH 7.5; 200 mM NaCl; 20 mM imidazole, detergent at 3x CMC.
Elution buffer: 50 mM HEPES, pH 7.5; 200 mM NaCl; 250 mM imidazole, detergent at 3xCMC
|
Column 1 Procedure: The solubilised membrane protein was batch bound to Cobalt Talon resin equilibrated with extraction buffer at 4°C for one hour and subsequently poured into a gravity flow glass column. The column was then washed with 30CV of wash buffer followed by elution in 1CV fractions until all the protein was eluted.
|
Column 2: Size Exclusion Chromatography. Superdex200 (GE healthcare) |
Column 2 Buffer:
Gel Filtration buffer: 20 mM HEPES pH 7.5, 200 mM NaCl,detergent at 1.2xCMC
|
Column 2 Procedure: The protein was concentrated and applied to a Superdex 200 gel filtration column equilibrated with gel filtration buffer using an AKTA purifier system.
|
Enzymatic Treatment: TEV protease (1:4, TEV:protein) was added overnight at 4°C to purified protein. The protease was removed and tag cleaved protein was collected by binding to Cobalt talon and collecting the flow-through. |
Mass spec characterization:
The purified protein was homogeneous and had an experimental mass of 55460Da, as expected from the primary sequence with a post translational modification of loss of methionine. Masses were determined by LC-MS, using an Agilent LC/MSD TOF system with reversed-phase HPLC coupled to electrospray ionisation and an orthogonal time-of-flight mass analyser. Proteins were desalted prior to mass spectrometry by rapid elution off a C3 column with a gradient of 5-95% methanol in water with 0.1% formic acid.
|
Protein Concentration: Protein was concentrated to ~20mg/ml using a 100kDa cut-off concentrator and back diluted to 9-11mg/ml. A three-fold excess of a farnesylated nonapeptide corresponding to the C-terminus of prelamin A (residues 656-664; QSPQNC(farn)SIM) was added prior to setting up crystallization plates |
Crystallization: P1 crystals were obtained under similar conditions to the native enzyme at 20°C. Crystals grew from sitting drops (150nl) comprising 100nl of concentrated protein / peptide complex and 50nl of reservoir equilibrated against 20µl of reservoir solution (0.1M HEPES pH 7.0, 0.1M calcium chloride, 29% (v/v) PEG 400). |
Data Collection:
Resolution: 3.8Å
Data were collected at 100K on the microfocus beamline I24 at Diamond Light Source using helical / straight line scans with a beamsize of 10µm x 10µm (wxh).
|
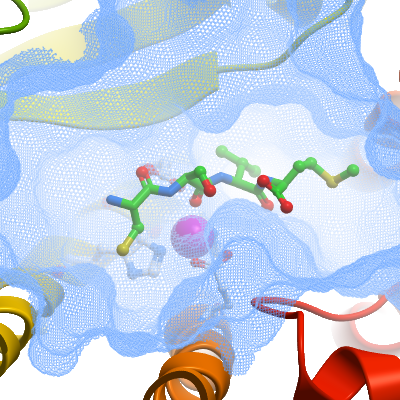
Structure Solution: Datasets collected from ZMPSTE24 / prelamin A peptide (Q656-M664) co-crystals showed that the peptide-binding site was empty. These co-crystals were then soaked with reservoir solution containing 5mM CSIM tetrapeptide and OGNG/CHS detergent for 24h at 6°C. The coordinates of the native P1 structure (4AW6) were used as a starting model for refinement with BUSTER. Clear difference Fo-Fc electron density in each of the four molecules in the asymmetric unit was interpreted based on the known binding mode of peptides and inhibitors to metalloproteases. The peptide register, with Ser662 and Ile663 positioned in the S1 and S1' subsites respectively, was determined based on both the length of the electron density, and the definition of the C-terminal. The entire tetrapeptide was modeled in two cases (molecules B&D) whereas only C661-I663 was modeled in molecules A & E. In all cases, the quality of the electron density was not sufficient to resolve the CD1 atom of Ile663 and this atom was not included in the model. Refinement was performed with BUSTER with TLS, NCS and LSSR restraints to the native structure. |