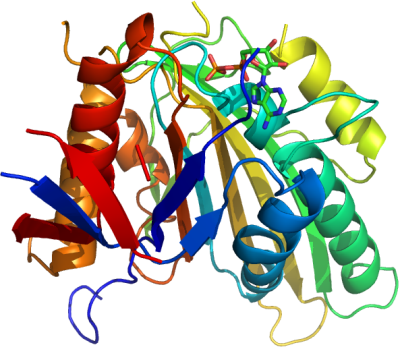
KIF22
PDB:3BFN
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:NP_015556
Entry Clone Source:MGC (AT50-E8)
SGC Clone Accession:
Tag:N-terminal hexahistidine tag
Host:E.coli. BL21 (DE3) codon(+) RIL
Construct
Prelude:
Sequence:mhhhhhhssgrenlyfqgPPARVRVAVRLRPFVDGTAGASDPPCVRGMDSCSLEIANWRNHQETLKYQFDAFYGERSTQQDIYAGSVQPILRHLLEGQNASVLAYGPTGAGKTHTMLGSPEQPGVIPRALMDLLQLTREEGAEGRPWALSVTMSYLEIYQEKVLDLLDPASGDLVIREDCRGNILIPGLSQKPISSFADFERHFLPASRNRTVGATRLNQRSSRSHAVLLVKVDQRERLAPFRQREGKLYLIDLAGSEDNRRTGNKGLRLKESGAINTSLFVLGKVVDALNQGLPRVPYRDSKLTRLLQDSLGGSAHSILIANIAPERRFYLDTVSALNFAARSKEVINRPFTNESLQPHALGPVKLSQKELLGPPEAK
Vector:p28a-mhl-tev
Growth
Medium:Terrific Broth medium
Antibiotics:
Procedure:We prepared the seeds by inoculating glycerol stock of E. coli cells BL21-CodonPlus (DE-3)-RIL containing the plasmid into 200 mL of Luria-Bertani medium. After overnight growth, all of the seeds were inoculated into 6 L of Terrific Broth medium in the presence of 50 µg/mL of kanamycin and 50 µg/mL chloramphenicol at 37 ºC and grown to an OD600 between 3-5. Cells were then induced by isopropyl-1-thio-D-galactopyranoside at the final concentration of 0.5 mM and grown overnight at 18 ºC in the SGC LEX bubbling system.
Purification
Procedure
Cultures were centrifuged and the cell pellets were harvested and stored at -80 ºC before use. Cells were thawed and suspended in 500 mL binding buffer (1X PBS, 0.5 M NaCl, 5 mM Imidazole, 5 mM BME, pH 7.0) with 0.5% (v/v) protease inhibitor cocktail (Sigma), 1600 units Benzonase (Sigma), and lysed with microfluidizer. The lysate was centrifuged at 16000 rpm for 60 min and the supernatant was used for subsequent steps of purification. All the extraction steps were carried out at 4 ºC.
Column 1: Ni-NTA beadsBuffers: Wash buffer 1: 1X PBS, 0.5 M NaCl, 30 mM Imidazole, 5 mM BME, pH 7.0. Elution buffer: 20 mM Hepes, 0.5 M NaCl, 300 mM Imidazole, pH 7.0, 5 mM BME, 2 mM MgCl2.Procedure: 1 ml of Ni-NTA suspension solution (50%) was added into 80 ml cell lysis supernatant solution. The mixture was shaken for 1 hour at 4 ºC. Beads were collected with centrifuge at 2500 rpm, 5 minutes. Each fraction of Beads was washed with 100 ml washing buffer, then collected with centrifuge. Protein was eluted with 15 ml elution buffer.
Column 2 : Size exclusion chromatography (Superdex 75 26/60)SEC-Buffers: 20 mM Hepes, pH 7.0, 500 mM NaCl, 1 mM TCEP. The fractions eluted of the Ni-affinity chromatography applied to a Superdex S75 column equilibrated in SEC buffer at a flow rate of 2.0 ml/min. Eluted fractions were 95% pure as judged by SDS-PAGE.
Extraction
Procedure
Cultures were centrifuged and the cell pellets were harvested and stored at -80 ºC before use. Cells were thawed and suspended in 500 mL binding buffer (1X PBS, 0.5 M NaCl, 5 mM Imidazole, 5 mM BME, pH 7.0) with 0.5% (v/v) protease inhibitor cocktail (Sigma), 1600 units Benzonase (Sigma), and lysed with microfluidizer. The lysate was centrifuged at 16000 rpm for 60 min and the supernatant was used for subsequent steps of purification. All the extraction steps were carried out at 4 ºC.
Concentration:32 mg/ml
Ligand
Mg(II), ADPMassSpec:
Crystallization:Crystals were obtained using the vapor diffusion method and a protein concentration of 32 mg/ml containing 5 molar fold of ADP and MgCl2. 2 µl of the concentrated protein mixed with 1% (w/w) Endoproteinase Glu-C V8 protease, then mixed with 2 µl of a well solution containing 3.2 M NaCl, 0.1 M Tris pH 8.5. Crystals appeared after one day at 18 °C.
NMR Spectroscopy:
Data Collection:Crystals were cryo-protected using ethylene glycol, and flash frozen in liquid nitrogen. Diffraction data were collected at APS to 2.3 Å.
Data Processing: