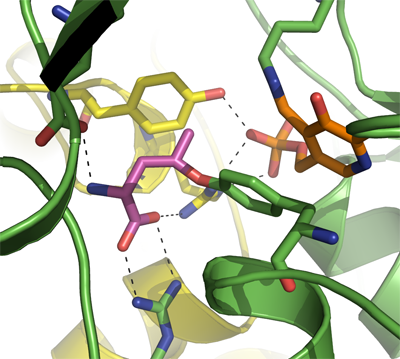
CTH
PDB:3COG
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:BC015807
Entry Clone Source:Mammalian Gene Collection
SGC Clone Accession:Tag:N-terminal hexahistidine tag with integrated TEV protease cleavage site: mhhhhhhssgvdlgtenlyfq*s(m).
Host:E.coli BL21(DE3) (Novagen)
Construct
Prelude:Sequence:mhhhhhhssgvdlgtenlyfq*smQEKDASSQGFLPHFQHFATQAIHVGQDPEQWTSRAVVPPISLSTTFKQGAPGQHSGFEYSRSGNPTRNCLEKAVAALDGAKYCLAFASGLAATVTITHLLKAGDQIICMDDVYGGTNRYFRQVASEFGLKISFVDCSKIKLLEAAITPETKLVWIETPTNPTQKVIDIEGCAHIVHKHGDIILVVDNTFMSPYFQRPLALGADISMYSATKYMNGHSDVVMGLVSVNCESLHNRLRFLQNSLGAVPSPIDCYLCNRGLKTLHVRMEKHFKNGMAVAQFLESNPWVEKVIYPGLPSHPQHELVKRQCTGCTGMVTFYIKGTLQHAEIFLKNLKLFTLAESLGGFESLAELPAIMTHASVLKNDRDVLGISDTLIRLSVGLEDEEDLLEDLDQALKAAHPPSG
Vector:pNIC-Bsa4
Growth
Medium:Antibiotics:Procedure:Cells from a glycerol stock were streaked onto an agar plate. Colonies were used to inoculate 2 x 20 ml TB supplemented with 8 g/l glycerol, 100 µg/ml kanamycin at 30 ºC overnight. The overnight culture (40 ml) was used to inoculate 2 x 1.5 l TB supplemented with 8 g/l glycerol, 50 µg/ml kanamycin and approximately 200 µl BREOX FMT 30 anti-foam solution (Cognis Performance Chemicals UK Ltd). The cultures were grown in a LEX bioreactor system (Harbinger Biotechnology) at 37 ºC until OD600 reached ~2. The bottles were down-tempered to 18 ºC over a period of 1 hour before target expression was induced by addition of 0.5 mM IPTG. Expression was allowed to continue overnight and cells were harvested the following morning by centrifugation (5,500 x
g, 10 min, 4 ºC). The resulting cell pellet (67.1 g wet cell weight) was resuspended in lysis buffer (2 ml/g cell pellet), supplemented with 1.5 tablets of Complete EDTA-free protease inhibitor (Roche Applied Science). The cell suspension was stored at -80 ºC.
Purification
ProcedureColumns
IMAC: Ni-charged 1 ml HiTrap Chelating HP (GE Healthcare)
Gel filtration column: HiLoad 16/60 Superdex 200 Prep Grade (GE Healthcare)
Procedure
The N-terminal histidine tag was proteolytically removed by incubating the target protein with His-tagged TEV protease at a molar ratio of 50:1 at 4 ºC for 72 hours. The proteolytic reaction went to completion, as judged by SDS-PAGE. Target protein was purified from tag and protease by passing the reaction mixture over a Ni-charged 1 ml HisTrap FF column (GE Healthcare) pre-equilibrated with IMAC wash1 buffer. The cleaved protein was concentrated and the buffer was changed to GF buffer containing 2 mM TCEP using an Amicon Ultra-15 centrifugal filter device with 10,000 NMWL (Millipore). The final protein concentration was determined to 24.5 mg/ml in a volume of 1 ml. The identity of the protein was confirmed by mass spectrometry.
Extraction
ProcedureThe cell suspension was quickly thawed in water and 4000 U Benzonase was added. Cells were disrupted by sonication (Vibra-Cell, Sonics) at 80% amplitude for 3 min effective time (pulsed 4s on, 4s off) and cell debris was removed by centrifugation (49,000 x
g, 20 min, 4 ºC). The supernatant was decanted and filtered through a 0.45 µm flask filter.
Concentration:LigandMassSpec:Crystallization:Crystals were obtained by the hanging drop vapour diffusion method in a 24-well plate containing 500 ml well solution. 1.5 µl of the protein sample (diluted to 10 mg/ml) was mixed with 1.5 µl of well solution consisting of 0.2 M ammonium citrate, pH 5.3 and 20% PEG 3350. The plate was streak-seeded with crystals obtained in an optimization screen where 0.1 µl protein (12.2 mg/ml) had been mixed with 0.2 µl well solution consisting of 0.2 M ammonium citrate, pH 5.3 and 30% PEG 3350 in a sitting drop at 20 ºC. The protein used for seeding was obtained in the same way as described above except that the his-tag had not been removed. The plate was incubated at 20 ºC and the crystal was soaked for 55 min in well solution supplemented with 10 mM DL-propargyl glycine. The crystal was then transferred to cryo solution consisting of well solution and 20% glycerol and flash-frozen in liquid nitrogen.
NMR Spectroscopy:Data Collection:Diffraction data to 2.0 Å resolution was collected at BESSY beamline ID-14.1.
Data Processing:The structure was solved by molecular replacement using the apo structure of cystathionine gamma lyase as template (PDB: 2NMP). The space group was P212121 with cell dimensions a=105.35 Å, b=107.22 Å, c=153.31 Å. Four monomers, forming a tetramer, were located in the asymmetric unit. Refmac5 was used for refinement and Coot for model building. Data in the interval 20.0 - 2.20 Å resolution was used and at the end of the refinement the R values were R=16.1% and Rfree=20.4%. Coordinates for the crystal structure were deposited in the Protein Data Bank with accession code 3COG.