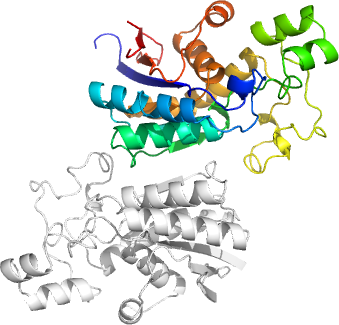
Cryptosporidium parvum phoshoglycerate mutase
PDB:3D8H
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:cgd7_4270
Entry Clone Source:SGC Clone Accession:CP-PF11_0208:H10
Tag:mgsshhhhhhssglvprgs
Host:BL21(DE3)-Codon-Plsus-RIL
Construct
Prelude:Sequence:TYKLTLIRHGESEWNKENRFTGWTDVSLSEQGVSEAIEAGRMLLEKGFKFDVVYTSVLKRAIMTTWTVLKELGNINCPIINHWRLNERHYGALQGLNKSETASKFGEDQVKIWRRSFDVPPPVLEKSDPRWPGNELIYKGICPSCLPTTECLKDTVERVKPYFEDVIAPSIMSGKSVLVSAHGNSLRALLYLLEGMTPEQILEVNIPTACPLVLELDDYLKVTKKYYLISEEELKAKMEAVANQGKAK
Vector:p28a-thrombin-lic
Growth
Medium:TB
Antibiotics:Procedure:Cryptosporidium parvum CP-PGAM1 (cgd7_4270) was expressed in E. coli BL21(DE3) CodonPlus-RIL in Terrific Broth (TB) in the presence of kanamycin/chloramphenicol (50 microgram/mL and 25 microgram/mL, respectively). A single colony was inoculated into 10 mL of LB with of kanamycin/chloramphenicol (50 microgram/mL and 25 microgram/mL respectively) in a 50 mL Falcon tube and incubated with shaking at 250 rpm overnight at 37 °C. The culture was transferred into 50 mL of TB with 50 microgram/mL kanamycin in a 250 mL shaking flask and incubated at 37 °C for 3 hours. Then the culture was transfer into 1.8 L of TB with 50 microgram/mL kanamycin and 0.3 mL of antifoam (Sigma) in a 2 L bottle and cultured using the LEX system to an OD600 > 5, cooled to 15 °C, and induced with 0.5 mM isopropyl-1-thio-D-galactopyranoside (IPTG) overnight at 15 °C.
Purification
ProcedureThe cleared lysate was loaded onto a column prepacked with 10 g DE52 (Whatman) anion exchange resin (previously activated with 2.5 M NaCl and equilibrated with Binding Buffer); and subsequently onto a 1.0 - 2.5 mL Ni-NTA (Qiagen) column pre-equilibrated with Binding Buffer at approximately 1 - 1.5 mL/min. The volume of the Ni-NTA resin was pre-determined by the predicted protein yield from test expression analysis. After the lysate was loaded, the DE52 was further washed with 20 mL of Binding Buffer. The Ni-NTA column was then washed with 200 mL of Wash Buffer at 2 - 2.5 mL/min. After washing, the protein was eluted with 15 mL of Elution Buffer. The eluted CP-PGAM1 (cgd7_4270) was dialysed against 10 mM HEPES, pH 7.5 and 500 mM NaCl then concentrated using a 15 mL Amicon Ultra centrifugal filter device (Millipore). The concentrated sample was flash frozen in N2(l) and stored at -80 °C. Stock concentration for CP-PGAM1 (cgd7_4270) is 15 mg/mL .
The His6-tag was cleaved with TEV overnight at 4 degC and dialysed into Crystal Buffer (10 mM HEPES, pH 7.5, 500 mM NaCl). The cleaved sample was loaded onto a Ni-NTA column and the flow through was collected and concentrated using a 15 mL Amicon Ultra centrifugal filter device (Millipore). The protein sample identity and purity were evalulated by mass spectroscopy and SDS-PAGE gel. The concentrated protein was stored at 4 degC.
Extraction
ProcedureThe culture was harvested by centrifugation. Pellets from 4 L of culture were resuspended to approximately 40 mL/L of cell culture in Binding Buffer with the addition of protease inhibitors (1 mM benzamidine and 1 mM phenylmethyl sulfonyl fluoride (PMSF)). Resuspended pellets stored at -80 degC were thawed overnight at 4 degC on the day before purification. Prior to mechanical lysis, each pellet from 1 L of culture was pretreated with 0.5 % CHAPS and 500 units of benzonase for 40 minutes at room temperature. Cells were mechanically lysed with a microfluidizer (Microfluidizer Processor, M-110EH) at approximately 18000 psi; and the cell lysate was centrifuged using a Beckman JA-25.50 rotor at ~75000 x g (24000 rpms) for 20 minutes at 10 degC.
Concentration:LigandMassSpec:Crystallization:The protein was crystallized at 20 degC in 25% PEG 3350, 0.2 M Ammonium acetate, 0.1 M Hepes, pH 7.5 using the hanging drop vapor diffusion method.
NMR Spectroscopy:Data Collection:Data Processing: