Entry Clone Source: Synthetic |
Entry Clone Accession: n/a |
SGC Construct ID: GSG2A-c009 |
GenBank GI number: gi|56790919 |
Vector: pNIC28-Bsa4. Details [PDF ]; Sequence [ FASTA ] or [ GenBank ] |
Tags and additions: Tag sequence: mhhhhhhssgvdlgtenlyfq*s(m) TEV-cleavable (*) N-terminal his6 tag. |
Final protein sequence:
mhhhhhhssgvdlgtenlyfq*sMGECSQ
KGPVPFSHCLPTEKLQRCEKIGEGVFGEV
FQTIADHTPVAIKIIAIEGPDLVNGSHQK
TFEEILPEIIISKELSLLSGEVCNRTEGF
IGLNSVHCVQGSYPPLLLKAWDHYNSTKG
SANDRPDFFKDDQLFIVLEFEFGGIDLEQ
MRTKLSSLATAKSILHQLTASLAVAEASL
RFEHRDLHWGNVLLKKTSLKKLHYTLNGK
SSTIPSCGLQVSIIDYTLSRLERDGIVVF
CDVSMDEDLFTGDGDYQFDIYRLMKKENN
NRWGEYHPYSNVLWLHYLTDKMLKQMTFK
TKCNTPAMKQIKRKIQEFHRTMLNFSSAT
DLLCQHSLFK |
Host: BL21(DE3)-R3-pRARE2 (previously known as Rosetta) |
Growth medium, induction protocol: 1ml from a 10 ml overnight culture containing 50 µg/ml kanamycin and 35 µg/ml chloramphenicol was used to inoculate 1 liter of LB media containing the same concentration of antibiotics. Cultures were grown at 37°C until the OD600 reached ~0.3. After that the temperature was adjusted to 20°C. Expression was induced over night using 1mM IPTG at an OD600 of 0.8. The cells were collected by centrifugation and the pellet resuspended in binding buffer and frozen.
Binding buffer: 50mM HEPES pH 7.5; 500 mM NaCl; 5% glycerol, 10 mM imidazole. |
Extraction method: Cell pellets were lysed by C5 high pressure homogenizer (Avestin). The lysate was centrifuged at 21,000 rpm for 60 minutes and the supernatant collected for purification. |
Column 1: Ni-affinity chromatography. |
Buffers: Binding buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 5% Glycero, 10mM Imidazole. Wash buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 30 mM Imidazole, 5% glycerol. Elution buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 50 to 250 mM imidazole, 5% Glycerol. |
Procedure: 5 ml of 50% Ni-NTA slurry (Qiagen) was applied to a 1.5 x 10 cm gravity column. The column was equilibrated with 50 ml binding buffer. The lysate was applied to the column which was subsequently washed with 50 ml wash buffer 1 and 2. The protein was eluted by gravity flow by applying 5 ml portions of elution buffer with increasing concentration of imidazole (50 mM, 100 mM, 150mM, 250 mM); fractions were collected until essentially all protein was eluted. The eluted protein was analyzed by SDS-PAGE. DTT was added to the protein sample to a final concentration of 5mM. |
Column 2: Size exclusion chromatography (Superdex S75, 60 x 1cm) |
SEC-Buffers: 50 mM Hepes, pH 7.5, 150 mM NaCl, 5 mM DTT. |
Procedure: The fractions eluted of the Ni-affinity chromatography were concentrated to about 4 mls using Centricon concentrators (10kDa cut off). The concentrated protein was applied to a Superdex S75 column equilibrated in SEC buffer at a flow rate of 0.8 ml/min. Eluted fractions were 95% pure as judged by SDS-PAGE. |
Protein concentration: Centricon with a 30 kDa cut off in SEC-buffer. |
Mass Spec Characterisation: The mass of the protein was calculated to be 40655 Da and experimentally determined mass was 40663 Da for the His tag containing protein. The identity of the protein was reconfirmed to be correct by DNA sequencing both DNA strands of this expression construct. |
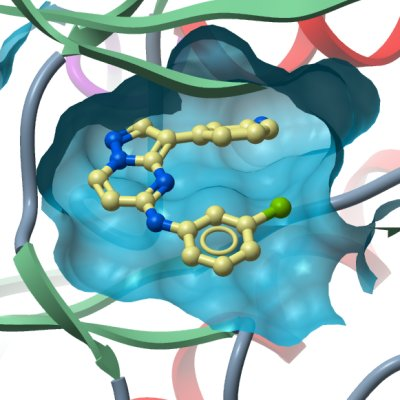
Crystallization: The protein was concentrated in gel filtration buffer to a protein concentration of 10.0 mg/ml. All crystallizations were carried out using sitting drop vapor diffusion at 4°C. Crystallization was performed by matrix-seeding with crushed native crystals into 200nl drops composed of equal volumes of native protein (10 mg/ml) with the ligand, 3-(3-aminophenyl)-N-(3-chlorophenyl)pyrazolo[1,5-α]pyrimidin-5-amine (1mM) and reservoir solution containing: 0.1M PCB pH 7.0; 30.0% PEG 1K. Crystals appeared within a few days |
Data Collection: Crystals were cryo-protected using the well solution supplemented with 25% Ethylene Glycol and flash frozen in liquid nitrogen. Diffraction data were collected from a single crystal on a Rigaku FR-E SuperBright at a single wavelength of 1.5Å. The structure was solved by molecular replacement and refined to 2.0 Å. |