Entry Clone Source: Ordered-synthetic |
Entry Clone Accession: n/a |
SGC Construct ID: CRYL1A-c014 |
GenBank GI number: gi|115430219 |
Vector: pNIC28-Bsa4. Details [PDF ]; Sequence [ FASTA ] or [ GenBank ] |
Tags and additions: TEV-cleavable (*), N-terminal histag. Tag sequence: mhhhhhhssgvdlgtenlyfq*sm |
Protein sequence (tag sequence in lowercase):
smAGCVVIVGSGVIGRSWAMLFASGGFQV
KLYDIEQQQIRNALENIRKEMKLLEQAGS
LKGSLSVEEQLSLISGCPNIQEAVEGAMH
IQECVPEDLELKKKIFAQLDSIIDDRVIL
SSSTSCLMPSKLFAGLVHVKQCIVAHPVN
PPYYIPLVELVPHPETAPTTVDRTHALMK
KIGQCPMRVQKEVAGFVLNRLQYAIISEA
WRLVEEGIVSPSDLDLVMSEGLGMRYAFI
GPLETMHLNAEGMLSYCDRYSEGIKHVLQ
TFGPIPEFSRATAEKVNQDMCMKVPDDPE
HLAARRQWRDECLMRLAKLKSQV |
Host: BL21(DE3)-R3-pRARE2 |
Growth medium, induction protocol: Medium: TB + 50 µg/ml Kanamycin + 34 µg/ml chloramp. 2 x1 liter TB in 3-L flasks were inoculated with 2 x10 ml overnight culture and grown at 37°C. The protein expression was induced with 0.1 mM IPTG at OD600 = 2.5 at 18°C overnight. The cells were collected by centrifugation and frozen at -80°C |
Extraction buffer, extraction method: Lysis buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 20 mM Imidazole, 5% glycerol, 0.5 mM TCEP, Complete® protease inhibitors (1 tablet/50 ml) and 5 U/ml of Benzonase. Cell pellet from 2 liter was resuspended in 100 ml binding buffer. The cells were disrupted by sonication and nucleic acids and cell debris removed by adding 0.15% PEI, followed by centrifugation for 40 minutes at 15 K rpm (JA17 rotor). |
Column 1: 4 ml Ni-Sepharose 6 Fast Flow |
Buffers: Binding buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 20 mM Imidazole, 5% glycerol, 0.5 mM TCEP and 5 U/ml of Benzonase; Wash and elution buffers: (a) 50 mM HEPES pH 7.5, 500 mM NaCl, 40 mM Imidazole, 5% glycerol; (b) 50 mM HEPES pH 7.5, 500 mM NaCl, 60 mM Imidazole, 5% glycerol; (c) 50 mM HEPES pH 7.5, 500 mM NaCl, 80 mM Imidazole, 5% glycerol; (d) 50 mM HEPES pH 7.5, 500 mM NaCl, 250 mM Imidazole, 5% glycerol. |
Procedure: The cell lysate was applied onto a 4 ml Ni-NTA column equilibrated with binding buffer. The column was subsequently washed with 20 ml of binding buffer and the protein was eluted using a stepwise gradient of imidazole. The eluted protein was collected and analyzed by SDS-PAGE. |
Column 2: Hiload 16/60 Superdex 200 prep grade 120 ml (GE/Amersham Biosciences) |
Buffers: 10 mM HEPES, pH 7.5, 500 mM NaCl, 5% glycerol |
Procedure: The fractions eluted from the Ni-affinity chromatography were concentrated to 3.5 ml and then incubated with 5mM DTT. The protein was filtered through a 0.22 mm filter and applied to a Superdex S200 column at a flow rate of 0.8 ml/min. The eluted proteins were collected in 1.8 ml fractions |
Enzymatic treatment: TEV cleavage. |
Column 3: Ni-NTA (TEV clean up) |
Buffer: 10 mM HEPES, pH 7.5, 500 mM NaCl, 5% glycerol, 0.5 mM TCEP |
Procedure: The Histidine-tag was cleaved with 150 µg of TEV protease per 10 mg protein at 4°C for 48 hours. |
TEV clean up: The TEV cleaved protein was applied to a 0.8 ml Ni-NTA column and the flow through collected. The column was washed with 5 ml buffer and the flow through and the wash fraction were analysed by SDS-PAGE. |
Concentration: The protein was concentrated in Amicon (3 K) to 20.5 mg/ml. The protein concentration was determined spectrophotometrically using the predicted molar extinction coefficient 26930 (M-1cm-1). |
Mass spectrometry characterization: ESI-MS revealed that the protein had the expected mass of 34 837 Da. |
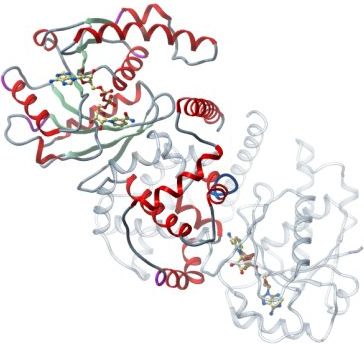
Crystallisation: CRYL1A was crystallised by vapor diffusion at 20°C from a sitting drop consisting of 100 nl protein (20.5 mg/l and 5 mM NAD) and 50 nl well solution. The drop was equilibrated against well solution containing 2 M ammonium sulfate. The crystal was transferred to a cryoprotectant composed of 25 % gycerol before flash-cooling in liquid nitrogen. |
Data Collection: Resolution: 2.0 Å; X-ray source: Synchrontron SLS-X10. |