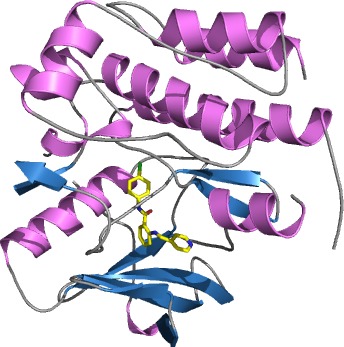
FLT1
PDB:3HNG
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:gi|24660372
Entry Clone Source:SGC Toronto. Synthetic gene
SGC Clone Accession:FLT1A-s002
Tag:N-terminal hexahistidine tag with integrated TEV protease cleavage site: mhhhhhhssgvdlgtenlyfq*sm
Host:E.coli BL21(DE3) R3 pRARE, where R3 denotes a derivative of BL21(DE3) resistant to a strain of T1 bacteriophage (SGC Oxford) and the pRARE plasmid originating from the Rosetta strain (Novagen) supplies tRNAs for rare codons.
Construct
Prelude:Sequence:mhhhhhhssgvdlgtenlyfqsmPDEVPLDEQCERLPYDASKWEFARERLKLGKSLGRGAFGKVVQASAFGIKKSPTCRTVAVKMLKEGATASEYKALMTELKILTHIGHHLNVVNLLGACTKQGGPLMVIVEYCKYGNLSNYLKSKRDLFFLNKDAALHMEPKKEKMEPGLEQGKKPRLDSVTSSESFASSGFQEDKSLSDVEEEEDSDGFYKEPITMEDLISYSFQVARGMEFLSSRKCIHRDLAARNILLSENNVVKICDFGLARDIYKNPDYVRKGDTRLPLKWMAPESIFDKIYSTKSDVWSYGVLLWEIFSLGGSPYPGVQMDEDFCSRLREGMRMRAPEYSTPEIYQIMLDCWHRDPKERPRFAELVEKLGDLL
Vector:pNIC-Bsa4
Growth
Medium:Antibiotics:Procedure:Cells from a glycerol stock were grown in 2 x 50 ml TB supplemented with 8 g/l glycerol, 100 µg/ml kanamycin and 34 µg/ml chloramphenicol at 37 °C overnight. The overnight culture (100 ml) was used to inoculate 9 l TB (divided into 6 x 1.5 l bottles) supplemented with 8 g/l glycerol, 50 µg/ml kanamycin and approximately 200 µl 204 Antifoam A6426 (Sigma) per bottle. The culture was grown in a LEX bioreactor system (Harbinger Biotechnology) at 37 °C until OD600nm had reached 1.6. The culture was down-tempered to 18 °C over a period of 1 hour before target expression was induced by addition of 0.5 mM IPTG. Expression was allowed to continue overnight and cells were harvested the following morning by centrifugation (4,400 x
g, 10 min, 4 °C). The resulting cell pellet (152 g wet cell weight) was then stored at -80 °C.
Purification
ProcedureColumnsIMAC: Ni-charged 5 ml HisTrap HP (GE Healthcare)
Gel filtration column: HiLoad 16/60 Superdex 75 Prep Grade (GE Healthcare)
Ion exchange column : MonoQ 5/50 GL (GE Healthcare)
Procedure
The whole purification procedure was performed at 4 ºC. Protein samples were loaded onto three separates IMAC columns previously equilibrated in Lysis buffer. Each IMAC column was then washed with, first, 100 ml of IMAC wash1 buffer followed by 100 ml of IMAC wash2 buffer. These operations were carried on using a peristaltic pump. The three IMAC columns were then connected in series on an ÄKTA Prime system (GE Healtchare). Bound protein was eluted from the IMAC columns with IMAC elution buffer then loaded onto the gel filtration column equilibrated in GF buffer. Fractions containing the target protein were pooled.
Tag removal and unphosphorylation
Proteolytic removal of the N-terminal histidine tag and unphosphorylation were performed in the same step. Protein sample was incubated with His-tagged TEV protease in a molar ratio of 10:1, 500 U of Calf Intestine Alkaline Phosphatase (CIAP) (Fermentas) and 10 mM MgCl2 at 4 ºC overnight under agitation. The proteolytic reaction went to completion, as judged by SDS-PAGE. The reaction mix was supplemented with 20mM imidazole and incubated with 3 ml Ni-NTA resin (Qiagen) equilibrated in TEV purification buffer. The sample was loaded onto an empty gravity flow column (Qiagen) and the flowthrough was collected.
Ion exchange
The sample was concentrated to 5 ml in a Vivaspin 10,000 MWCO (Sartorius) then diluted to 20 ml in MonoQ low salt 7.5 buffer. This step was done twice. The protein was loaded onto the ion exchange column equilibrated in MonoQ low salt pH 8.5 buffer. A gradient between MonoQ low salt pH 8.5 buffer and MonoQ high salt pH 8.5 buffer was applied onto the column. Fractions eluted between 7 and 9% of MonoQ high salt pH 8.5 buffer were pooled, supplemented with 200 mM NaCl and concentrated to 14mg/ml in a Vivaspin 10,000 MWCO (Sartorius). Identity and level of phosphorylation of FLT1 were assessed by mass spectrometry.
Extraction
ProcedureThe pellet was splitted into three equal parts. Each part was resuspended into a final volume of 150ml of Lysis buffer (including the volume of the pellet) supplemented with a knife edge of lysozyme (Fluka), one tablet of Complete EDTA-free protease inhibitor (Roche Applied Science) and 1000 U Benzonase (Merck). The samples are left under agitation for 1 h at 4 C. Cells were disrupted by sonication (Vibra-Cell, Sonics) at 80% amplitude for 6 min effective time (pulsed 4s on, 4s off) and cell debris was removed by centrifugation (49,000 x
g, 40 min, 4 ºC). The supernatant was decanted and filtered through a 0.45 µm flask filter.
Concentration:LigandMassSpec:Crystallization:Protein sample was incubated for 30 min at room temperature with 2mM N-(4-Chlorophenyl)-2-((pyridin-4-ylmethyl)amino)benzamide (Calbiochem) dissolved in DMSO. The sample was then spinned for 30 min at 20,000 x
g in a table-top centrifuge. Supernatant was used to set-up crystallization plates. Crystals were obtained by the sitting drop vapour diffusion method in a 96-well plate. 0.1 µl protein solution (11.2 mg/ml) was mixed with 0.2 µl of well solution consisting of 0.2 M potassium formate and 20% PEG 3350. The plate was incubated at 4 ºC and 230 µm long thin plates appeared between 14 and 28 days. The crystals were quickly transferred to a cryo solution consisting of 0.2 M potassium formate, 22% PEG 3350, 0.3 M NaCl, 30% ethylene-glycol and flash frozen in liquid nitrogen.
NMR Spectroscopy:Data Collection:Diffraction data to 2.7 Å resolution was collected at ESRF beamline ID14-2.
Data Processing:The structure was solved by molecular replacement using the structure of human VEGFR2 kinase domain as template (PDB: 2QU5). The space group was I222 with cell dimensions a=58.04 Å, b=71.15 Å c=196.02 Å. The structure was solved using the Phenix suite of program. One monomer was located in the asymmetric unit. Iterative cycles of manual improvement and refinement were made respectively in Coot and Refmac. Data in the interval 44.99-2.7 Å resolution was used and at the end of the refinement the R values were: R=19.7% and Rfree=26.0%. Coordinates for the crystal structure were deposited in the Protein Data Bank, accession code 3HNG.