Entry Clone Source: TKC |
Entry Clone Accession: n/a |
SGC Construct ID: PIM1A-c001 |
GenBank GI number: gi|4505811 |
| Vector: pLIC- SGC1. Details [PDF]; Sequence [FASTA] or [GenBank |
Entry clone accession/ sequence: The expressed protein has the sequence from gi 33304198 which differs from gi 4505811 by a single change R250G (bold and red in the sequence below) |
Amplified construct sequence:
TACTTCCAATCCATGCAGAGCAGTAAGCGC
ATGCACCATCATCATCATCATTCTTCTGGT
GTAGATCTGGGTACCGAGAACCTGTACTTC
CAATCCATATGCTCTTGTCCAAAATCAACT
CGCTTGCCCACCTGCGCGCCGCGCCCTGCA
ACGACCTGCACGCCACCAAGCTGGCGCCCG
GCAAGGAGAAGGAGCCCCTGGAGTCGCAGT
ACCAGGTGGGCCCGCTACTGGGCAGCGGCG
GCTTCGGCTCGGTCTACTCAGGCATCCGCG
TCTCCGACAACTTGCCGGTGGCCATCAAAC
ACGTGGAGAAGGACCGGATTTCCGACTGGG
GAGAGCTGCCTAATGGCACTCGAGTGCCCA
TGGAAGTGGTCCTGCTGAAGAAGGTGAGCT
CGGGTTTCTCCGGCGTCATTAGGCTCCTGG
ACTGGTTCGAGAGGCCCGACAGTTTCGTCC
TGATCCTGGAGAGGCCCGAGCCGGTGCAAG
ATCTCTTCGACTTCATCACGGAAAGGGGAG
CCCTGCAAGAGGAGCTCGCCCGCAGCTTCT
TCTGGCAGGTGCTGGAGGCCGTGCGGCACT
GCCACAACTGCGGGGTGCTCCACCGCGACA
TCAAGGACGAAAACATCCTTATCGACCTCA
ATCGCGGCGAGCTCAAGCTCATCGACTTCG
GGTCGGGGGCGCTGCTCAAAGACACCGTCT
ACACGGACTTCGATGGGACCCGAGTGTATA
GCCCTCCAGAGTGGATCCGCTACCATCGCT
ACCATGGCAGGTCGGCGGCAGTCTGGTCCC
TGGGGATCCTGCTGTATGATATGGTGTGTG
GAGATATTCCTTTCGAGCATGACGAAGAGA
TCATCAGGGGCCAGGTTTTCTTCAGGCAGA
GGGTCTCTTCAGAATGTCAGCATCTCATTA
GATGGTGCTTGGCCCTGAGACCATCAGATA
GGCCAACCTTCGAAGAAATCCAGAACCATC
CATGGATGCAAGATGTTCTCCTGCCCCAGG
AAACTGCTGAGATCCACCTCCACAGCCTGT
CGCCGGGGCCCAGCTAAAGTAAAGGTGGAT
AC
|
Final protein sequence (tag sequence in lowercase):
mhhhhhhssgvdlgtenlyfqsMLLSKINS
LAHLRAAPCNDLHATKLAPGKEKEPLESQY
QVGPLLGSGGFGSVYSGIRVSDNLPVAIKH
VEKDRISDWGELPNGTRVPMEVVLLKKVSS
GFSGVIRLLDWFERPDSFVLILERPEPVQD
LFDFITERGALQEELARSFFWQVLEAVRHC
HNCGVLHRDIKDENILIDLNRGELKLIDFG
SGALLKDTVYTDFDGTRVYSPPEWIRYHRY
HGRSAAVWSLGILLYDMVCGDIPFEHDEEI
IGGQVFFRQRVSSECQHLIRWCLALRPSDR
PTFEEIQNHPWMQDVLLPQETAEIHLHSLS
PGPS |
| Tags and additions: mhhhhhhssgvdlgtenlyfq*s(m). N-terminal his6 tag, TEV-protease cleavable (*). |
| Host: BL21(DE3) |
Growth medium, induction protocol: 5 ml overnight cultures in LB, 100 µg/ml ampicillin were grown at 37°C and 0.5 ml used to inoculate 1 litre of LB medium containing 100 µg/ml ampicillin. Cultures were grown at 37°C until they reached an OD600 of 1.2 and then cooled to 18°C over 20 minutes. Expression was then induced overnight using 0.5 mM IPTG. The cells were collected by centrifugation, transferred to 50 ml tubes, resuspended in 30 ml binding buffer, and frozen. Binding buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 5% Glycerol. |
| Extraction buffer, extraction method: The frozen cells were thawed on ice and 0.5mM TCEP and 1 mM PMSF added. Cells were lysed using a high pressure cell disruptor. The lysate was centrifuged at 17,000 RPM for 30 minutes. Supernatant was collected and binding buffer was added to 50 ml. |
Column 1: Ion exchange - Nucleic acid removal. DEAE cellulose (DE52, Whatmann), 10 gr of resin in 2.5 x 20 cm column. The resin was hydrated in 2.5M NaCl, then washed with 20 ml binding buffer prior to loading the sample. |
Buffers: Binding buffer |
| Column 1 Procedure: Supernatant was applied at gravity flow, followed by a wash with 40 ml binding buffer. The column flow-through was collected. |
Column 2: Ni-affinity. Ni-sepharose (Amersham), 5 ml of 50% slurry in 1.5 x 10 cm column, equilibrated with binding buffer. |
| Column 2 Buffers: Binding buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 5% Glycerol. Wash buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 20 mM Imidazole, 5% Glycerol. Elution buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 50 to 250 mM Imidazole, 5% Glycerol. |
| Column 2 Procedure: The flowthrough from column 1 was loaded by gravity flow on the Ni-sepharose column. The column was then washed with 50 ml wash buffer under gravity flow. The protein was eluted by applying 5-ml portions of elution buffer with increasing concentration of imidazole (50 mM, 100 mM, 250 mM); fractions were collected until essentially all protein was eluted. After elution DTT was added to a final concentration of 10 mM. |
| Enzymatic treatment : Dephosphorylation (a GST fusion with the lambda phosphatase) and TEV protease cleavage. Samples containing Pim1 were pooled and treated with lambda phosphatase and TEV protease overnight at 4°C. Protein was kept in elution buffer with the addition of 10 mM DTT and 0.05 mM MnCl2 (higher MnCl2 concentrations caused precipitation). |
Column 3: HiLoad 16/60 Superdex 200 gel filtration |
Column 3 Buffers: 50 mM Hepes pH 7.5, 250 mM NaCl |
| Column 3 Procedure: Dephosphorylated Pim1 protein was concentrated to 3 ml and ran on a S200 gel filtration column collecting 1.8 ml fractions. 10 mM DTT was added to the eluted protein for overnight storage |
Column 4: Ion exchange Mono Q column. |
Column 4 Buffers: A : 50 mM Hepes pH 7.5. B : 50 mM Hepes pH 7.5, 1000 mM NaCl. |
| Column 4 Procedure: Pim1 was applied to MonoQ in buffer A and eluted from the column by a linear gradient. |
| Concentration: Pim1 fractions containing dephosphorylated protein were pooled and concentrated in Centricons (10 kDa cut off). Phosphorylation state was confirmed using LC- ESI MS-Tof. |
| Mass spectrometry characterization: The purified protein was homogeneous and had an experimental mass of 35625 Da as expected from its primary structure and a single phosphorylation. Masses were determined by LC-MS, using an Agilent LC/MSD TOF system with reversed-phase HPLC coupled to electrospray ionisation and an orthogonal time-of-flight mass analyser. Proteins were desalted prior to mass spectrometry by rapid elution off a C3 column with a gradient of 5-95% acetonitrile in water with 0.1% formic acid. |
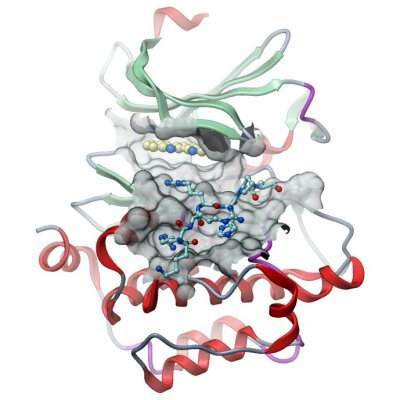
| Crystallization: Crystallization was carried out using the sitting drop vapor diffusion method at 4 °C. A 6 mL solution with 1 mg of PIM1 buffered in 50mMHepespH7.5, 250mMNaCl, 10mM DTTwas prepared containing 2 µL of inhibitor (from a 50 mM stock in DMSO) and 20 µL of a 5 mM solution of consensus peptide (ARKRRRHPSGPPTA-amide). This sample was then concentrated to 100 µL to yield a 10 mg/mL crystal sample. Crystals of the complex were grown by mixing 150 nL of the protein (5 mg/mL) with an equal volume of reservoir solution containing 0.2MNaF, 100mMbistrispropane pH 8.5, 20% PEG3350, and 10% ethylene glycol. Crystals grew to diffracting quality within a few days. Crystals were cryoprotected using the well solution supplemented with additional ethylene glycol and were flash frozen in liquid nitrogen. |
Data Collection: Resolution: 2.35 Å, X-ray source: rotating anode (Rigaku FR-E SuperBright), single wavelength 1.5 Å. |