Myristoylation is an irreversible, co-translational (during translation) protein modification found in animals, plants, fungi and viruses. It is an acylation process absolutely specific to the N-terminal glycine in proteins. In this reaction, a myristoyl group (a 14 carbon fatty acid) is covalently attached via an amide bond to the alpha-amino group of an N-terminal amino acid of a nascent polypeptide. Myristoylation plays a vital role in membrane targeting and signal transduction and is only the first step of the mechanisms by which a protein associates with the membrane[1].
The enzyme that carries out such a reaction is myristoyl-CoA:protein N-myristoyltransferace (NMT). NMT has been proved to be druggable target for the treatment of human African trypanosomiasis and other parasitic diseases[2]. To obtain a high resolution 3-D structure of human NMT is one of the critical steps in order to assist in the discovery of selective inhibitors for parasitic diseases.
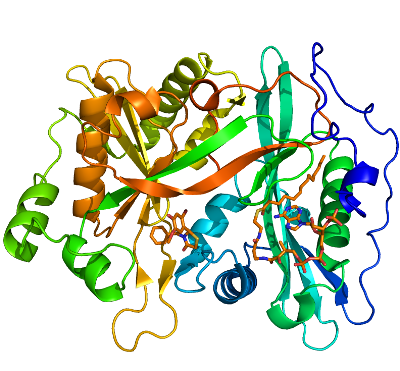
We determined a series of human NMT1 (HsNMT1) structures in complex with myristoyl-CoA [3IU1], and several lead compounds discovered by University of Dundee: DDD90096 (2-{4-Hydroxy-5-methoxy-2-[3-(4-methyl-piperazin-1-yl)-propyl]-phenyl}-3-pyridin-3-yl-thiazolidin-4-one) [3IU2], DDD86546 (2,6-Dichloro-4-(2-piperazin-1-yl-pyridin-4-yl)-N-(1,3,5-trimethyl-1H-pyrazol-4-yl)-benzenesulfonamide) [3IWE] and DDD90055 (3-Benzyl-2-(2-bromo-4-hydroxy-5-methoxy-phenyl)-thiazolidin-4-one) [3JTK]. The overall structure of HsNMT1 consists of a saddle-shaped mixed b-sheet and surrounded by several a-helices. Similar to C. albicans and S. cerevisiae NMT structures (PDB ID: 1NMT and 2NMT, respectively), HsNMT1 has a pseudo two fold symmetry with N-terminal half binds to myristoyl Co-enzyme A (M-CoA) and C-terminal half binds with peptide substrate. For M-CoA binding pocket, most residues forming the pocket are conserved and M-CoA molecule adapts similar conformation as seen in other available structures (1IID, 2NMT and 3H5Z). Larger deviation and conformational changes can be seen at peptide substrate binding site. For example, in HsNMT1+M-CoA+DDD85646 complex structure, the inhibitor DDD85646 occupied approximately the first five residues position of a peptide substrate (GLYASKLA) from S. cerevisiae NMT ternary complex structure ((1IID). However, the inhibitor adapts a different binding conformation as compared to the peptide substrate resulting a large movement of a loop region (Ab loop from S. cerevisiae NMT structure)[3]. The biggest shift in Ab loop is around 7A of Asp184 (Asp 106 in (1IID). The first pyridine ring from inhibitor locates in the approximate the Glycine residue position of the peptide substrate, with 4-NH group forming 4 potential hydrogen bonds with surrounding protein residues (two direct hydrogen bonds with the C-terminal end Oxt atom from Gln496 and Oδ1 of Asn246; two water mediated hydrogen bonding with Oη atoms of Tyr180 and Tyr192). The NH atom from second pyridine ring also form a hydrogen The 2-Cl atom from inhibitor DDD85646 makes van-der-val (VDW) contact with the side chain of a highly conserved Tyr420 and the 6-Cl atom forms another VDW contact with Gly472. These two VDW contacts together with forward mentioned 4 hydrogen bonds lock the position of the first three rings in the substrate binding pocket. The sulfonamide group from the inhibitor forms two hydrogen bonds with surrounding amino acids (one direct hydrogen bond with a highly conserved residue His298 and one water mediated hydrogen bond with a backbone NH group of Asp471), which create a significant bend of the pyrozole ring. The side chains of Val 181, Phe188, Phe190, Phe311 and Leu416 form a hydrophobic pocket for pyrozole ring while the 4-NH group from it forms a direct hydrogen bond with Oγ of Ser405, which further stabilizes the position of pyrozole ring.
The HsNMT1 structure provides important information about improving the selectivity of the inhibitor. Also with the empty space in peptide substrate position 6-8, there will be room to further design new generation of inhibitor.